Location: Home >> Detail
TOTAL VIEWS
Crop Breed Genet Genom. 2026;8(2):e260014. https://doi.org/10.20900/cbgg20260014
1 Syngenta Crop Protection, LLC, 9 Davis Drive, Durham, NC 27713, USA
2 Retired from Syngenta Crop Protection, LLC
* Correspondence: Craig Schuller, Qingli Liu
The introduction of genetic diversity from wild relatives to domesticated crops has been a successful practice to improve disease resistance and stress tolerance. There remain several challenges to this practice, including reproductive incompatibility, hybrid sterility, and repressed recombination between homoeologous chromosomes. In soybean, the need to diversify genetic pools is particularly compelling due to the narrow genetic base of commercial germplasm and the limited number of non-native landrace germplasm donors. Soybean perennial wild relatives (Glycine subgenus Glycine) contain resistance to major soybean diseases such as soybean rust and cyst nematode. These natural sources of genetic resistance remain largely untapped due to the difficulty in obtaining fertile progeny from wide crosses. We report a novel hybridization strategy utilizing auto-tetraploid soybean as the maternal parent in wide crosses. Combined with exogenous auxin applications, we demonstrated the robustness of this method by generating viable F1 progeny across tetraploid and diploid perennial accessions. The introgression of the soybean rust (SBR) resistance locus was tracked through successive backcross generations. Finally, fluorescence in situ hybridization (FISH) confirmed integration of the locus into a soybean chromosome, while subsequent evaluations established the stable inheritance and phenotypic expression of SBR resistance. These results demonstrate that balancing genome size prior to initial wide crosses can overcome barriers to hybridization. This finding offers a practical solution for breeders to broaden the genetic base of cultivated crops by facilitating the introgression of traits from previously inaccessible wild germplasm.
F1D, amphidiploid; BC, backcross; SBR, soybean rust; SCN, soybean cyst nematode; EBN, endosperm balance number; MTT, Thiazolyl Blue Tetrazolium Bromide; FISH, Fluorescence in situ hybridization; DAPI, 4’,6-diamidino-2-phenylindole; BAP, 6-Benzylaminopurine; IBA, Indole-3-butyric acid; Dicamba, 3,6-dichloro-0-anisic acid; ER, embryo rescue
Soybean (Glycine max) is one of the most important agricultural crops, serving as a major source of food, animal feed, protein, oil, and other products [1,2]. Efforts to improve soybeans through conventional breeding have been confined to the annual species Glycine max and G. soja. The 26 perennial wild relatives [1,3] have many agronomically useful traits, such as disease resistance. For example, accessions of G. argyrea, G. canescens, and G. tomentella were reported to confer resistance to pathogens such as soybean rust [4–8] and soybean cyst nematode [2,9,10].
The observed taxonomic complexity of perennial species makes performing intersubgeneric hybridization difficult, especially in the recovery of fertile progeny containing desired traits [3,8,11–14]. For example, the G. tomentella complex contains diploids (2n = 38 or 40) and allotetraploids (T1, T2, T3, T4, T5, and T6 with 2n = 78 or 80). These tetraploid species are composed of DDEE (T1), DDAA (T2 and T3), AAEE (T5), DDHH (T4), and EEHH (T6) subgenomes. Successful wide crosses producing an amphidiploid and backcross generation 1 (BC1) plants have been reported for two G. tomentella T1 lines, PI483218 and PI441001 [8,15]. Backcross-derived fertile lines were generated with subsequent individuals found to be susceptible to soybean rust, although the amphidiploid clones were resistant [8]. Other efforts to breed traits from perennial Glycine into soybean have not been successful at the F1 generation or the amphidiploid generation [3,8].
In this study, we modified the method of Singh and Nelson [8] outlined in Figure 1A (herein referred to as the modified Singh method). In the modified Singh method, approximately two weeks after pollination, immature F1 embryos were rescued and treated with colchicine for genome doubling. Two to three months later, tissue cultured F1D shoots were recovered directly without embryogenesis. These modifications fit into tissue culture operations in wide crosses with equivalent efficiency in generating BC1s using G. tomentella T1 PI441001 as reported in previous studies [8,16–18].
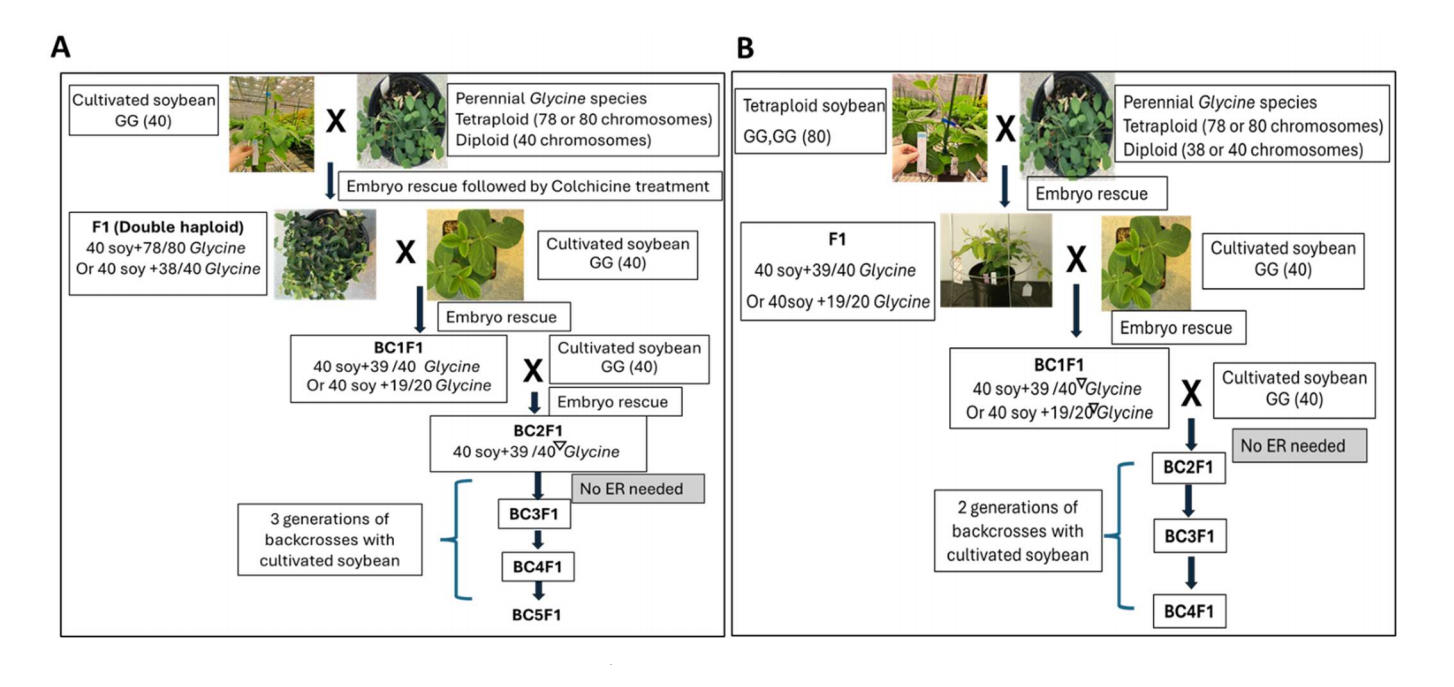 Figure 1. Comparison of workflows in (A) using soybean (modified from Singh and Nelson [9]) and (B) using genome doubled soybean (auto-tetraploid) as the female parent to conduct wide crosses with perennial Glycine species. (B) is referred to as the tetraploid method. Key differences are (1) No colchicine is used throughout the process in the tetraploid method; (2) Chromosome loss starts in the first backcross at F1 generation in tetraploid soybean method; (3) Embryo rescue (ER) is only needed to generate BC1F1 plantlets in the tetraploid method, while ER is required to generate BC2F1 plants in the modified Singh method; (4) the BC4 generation in the tetraploid method is equivalent to BC5 in the modified Singh method. Hence, one generation is saved in the tetraploid method. Inverted triangles indicate random chromosome losses.
Figure 1. Comparison of workflows in (A) using soybean (modified from Singh and Nelson [9]) and (B) using genome doubled soybean (auto-tetraploid) as the female parent to conduct wide crosses with perennial Glycine species. (B) is referred to as the tetraploid method. Key differences are (1) No colchicine is used throughout the process in the tetraploid method; (2) Chromosome loss starts in the first backcross at F1 generation in tetraploid soybean method; (3) Embryo rescue (ER) is only needed to generate BC1F1 plantlets in the tetraploid method, while ER is required to generate BC2F1 plants in the modified Singh method; (4) the BC4 generation in the tetraploid method is equivalent to BC5 in the modified Singh method. Hence, one generation is saved in the tetraploid method. Inverted triangles indicate random chromosome losses.
Using the modified Singh method, F1 amphidiploids were regenerated with several other G. tomentella accessions in the T1 genome type (PI583970, PI441001, PI483224, PI441008, and PI499939), as well as non-T1 genome types PI446958 (T2), and PI604489 (T4). However, producing genome-doubled F1s (F1Ds) in most cases was difficult due to the toxicity of the required colchicine treatment and the genetic incompatibility between soybean and perennial Glycine spp. Production of backcross 1 generation (BC1F1) plants was challenging due to extremely low fertility of the F1D. We were only successful in obtaining limited BC1F1 embryos with PI441001 and PI483224.
In previous studies in which BC1+ materials were generated, diploid soybean (2n = 40) was crossed to tetraploid G. tomentella accessions (2n = 78 or 80) [8,15]. Ploidy differences alone can lead to crossing failure due to a triploid block leading to the failure of endosperm development and nonviable progeny [19]. In soybean, cross-pollination between diploids and auto-tetraploids was unsuccessful in producing triploids [20], which could be explained by the endosperm balance number (EBN) hypothesis where the maternal:paternal genome ratio conforms to 2m:1p in the endosperm [21–23]. The mechanism of the EBN may be genomic imprinting, a transgenerational epigenetic regulation of alleles leading to parental expression biases [24]. Crosses in which the female has higher ploidy can fail due to premature endosperm cellularization, while the reciprocal cross can fail due to delayed endosperm cellularization [24]. When diploid female Arabidopsis thaliana was crossed to a tetraploid male, increased auxin synthesis and transport were observed in association with delayed endosperm cellularization and embryo failure [25]. Crosses where the female has the higher ploidy might benefit from the application of exogenous auxin to delay endosperm cellularization. We hypothesized that using the tetraploid soybean as a maternal donor would improve the success rate of wide crosses between soybean and perennial Glycine species facilitated by exogenous auxin application to potentially prevent early endosperm cellularization.
Previously generated tetraploid soybeans were shown to have lower self-fertility than their progenitors, likely due to abnormal meiotic chromosome behavior giving rise to improperly formed sterile pollen grains and pods [26,27]. The concept of using genome-doubled soybean (auto-tetraploid soybean) as described in Figure 1B (herein referred to as tetraploid soybean) is appealing in that the use of colchicine is skipped and saves one generation compared to the modified Singh method. We crossed tetraploid soybeans with G. tomentella T1 PI441001 to produce F1 hybrids with a more balanced chromosome complement GGDE (with GG referring to genome-doubled G. max and the DE genome parental progenitors) without using colchicine. The genome composition of an F1 derived from this method is identical to a BC1F1 derived from the modified Singh method as confirmed by fluorescence in situ hybridization. Then we successfully applied this tetraploid soybean method to additional tetraploid and diploid perennial accessions. Soybean rust resistance loci were successfully transferred and remained heritable across subsequent backcross generations, thereby establishing the use of tetraploid soybean as a viable strategy to broaden the genetic base of cultivated soybean.
Seven perennial Glycine accessions were selected to represent four different genome groups with a mix of diploids and tetraploids (Table 1), including T1, T2, and T5 tetraploids and a diploid (D-genome) [11,13]. Line selection was also based on reported successes in wide crosses or known resistance to soybean rust (SBR) or soybean cyst nematodes (SCN) as reported previously [7,9,10,28,29].
Five Syngenta soybean lines (HI2458, SJ0225, SJ1445, CC0653, and HI0211) were selected to test crossability and embryo recovery rates with the selected perennial Glycine lines. Lines SJ0225 and CC0653, which showed tolerance to greenhouse pests and diseases, were chosen for tetraploid generation using the following procedure. Embryos 1–3 mm in length were isolated on embryo rescue (ER) media containing (per L): 3.1 g Gamborg’s B5 basal salt, 1× Gamborg’s B5 vitamins, 40 g sucrose, 0.25 g casein hydrolysate enzymatic, 0.25 mg BAP, 0.75 g magnesium chloride hexahydrate, 0.5 g glutamine, 0.1 g serine; 0.1 g asparagine, 0.05 mg IBA, and 2 g Gelzan with 0.25–0.5 g colchicine for approximately three days. Embryos were then transferred to fresh ER media without colchicine. After approximately three weeks, embryos were transferred to maturation media (GSM) containing (per L): 3.2 g Schenck and Hilderbrandt basal salt mixture; 1 g myo-inositol; 5 mg thiamine; 5 mg nicotinic acid; 0.5 mg pyridoxine; 10 g sucrose; and 7.5 g purified agar until the formation of rooted shoots. Ploidy was confirmed using the CyStain UV Precise P flow cytometry kit and Sysmex/Partec Ploidy Analyzer (Sysmex Partec GmbH, Görlitz, Germany). Tetraploid plants were allowed to self-pollinate, and ploidy of the next generation was confirmed. Seed stocks were obtained after multiple generations of self-pollination.
Samples for pollen viability staining were prepared by collecting anthers from four flowers into 20 µL of MTT (Thiazolyl Blue Tetrazolium Bromide) staining solution containing 0.9 g MTT, 5 g sucrose, and 100 mL water in a microcentrifuge tube. A plastic pestle was used to release pollen grains from the anthers, and 10 µL was pipetted into the chamber of a disposable hemacytometer. Images were taken with an Olympus EP50 camera connected to an Olympus SZX7 microscope. Viable pollen grains were stained dark blue due to the reduction of the yellow MTT dye by mitochondrial reductases to form the product MTT-formazan.
Pollination and Dicamba Treatment in Production of F1s and BackcrossesWide crosses between tetraploid soybean (maternal) and tetraploid and diploid accessions of G. tomentella (paternal) were performed by brushing freshly collected pollen onto mature stigmas prior to anthesis. No emasculation was required. F1s were distinguished from self-pollinated plants by their smaller embryo size approximately 14 days after pollination, later by TaqMan assays in shoot stage and by FISH (Fluorescence in situ hybridization) using soybean centromere specific probes.
Dicamba (3,6-dichloro-0-anisic acid, C8H6Cl2O3), a synthetic auxin herbicide which is readily available as a weed control, was prepared as a 1 mg/mL stock solution and diluted to working concentrations ranging from 3–20 mg/L. The working solution was applied by lightly coating the pollination site using an atomizer, with two or more applications on non-consecutive days necessary to allow for F1 embryo development. Spray drift was minimized by holding a paper towel behind the pollination site during application. The first dicamba application was performed one to two days after pollen transfer, with the second application five to seven days after pollination, and the third application eight or nine days after pollination.
Pods containing F1 embryos were harvested approximately two weeks after pollination. Pods were briefly surface sterilized with 70% ethanol, followed by 10% Clorox bleach for 30 min. Sterilized pods were opened in a laminar flow hood with sterile tweezers. Immature F1 embryos were extracted onto ER medium. Approximately three weeks later, embryos were transferred to GSM medium until rooted shoots were ready for transfer to soil, usually in about two months.
F1 plants were backcrossed with diploid soybean (G. max) plants by brushing freshly collected pollen onto the mature F1 stigmas. Since F1s lack self-fertility, pollination can be performed on fully opened F1 flowers without the risk of self-pollination. Pods containing BC1 embryos were harvested approximately two weeks after pollination and sterilized, then embryos were extracted and carried through tissue culture as described above. In some cases, pods were left on F1 plants until small mature seeds were obtained. BC1F1 plants were backcrossed to diploid soybeans in the same way as the F1s. BC2F1 seeds were grown to maturity prior to harvesting.
Hybrid CharacterizationsFluorescence in situ hybridization (FISH) was used to distinguish between subgenome chromosomes in young root tips from the putative F1 plants using the method described by Gill et al. (2009) [30]. Two subfamilies of high-copy centromeric satellite repeats, CentGm-1 and CentGm-2, were used as probes because they only hybridize to genomic DNA of Glycine soja and G. max, and not to other Glycine species [30]. Probes were amplified with primers listed in Table S1 and labelled with Alexa Fluor® 488 by nick translation. All chromosomes were visualized using chromosome counterstain DAPI (4’,6-diamidino-2-phenylindole).
Root tip cytology was performed on multiple putative F1s to confirm the expected chromosome composition. Roots were first arrested in 0.05% 8-hydroxyquinoline for five hours and fixed in 90% acetic acid. The meristems of the root tips were briefly digested with 0.5% pectolyase and 1% cellulase to remove cell walls. A confocal image using the chromosome counterstain DAPI at 405 nm excitation was obtained using a Leica confocal laser scanning microscope LSM880 from Zeiss (Oberkochen, Germany) and using C-apochromat 40× or 63/×1.2 W Korr FCS M27 water-immersion objectives and the ZENblack software by Zeiss. Labelled centromeres of soybean chromosomes in the F1 hybrid were obtained using Alexa Fluor® 488 at 488 nm excitation showing green fluorescence. Confocal images were merged to distinguish soybean chromosomes from G. tomentella chromosomes with all soybean chromosomes stained with centromere probes in green fluorescence, while G. tomentella chromosomes were solely stained with DAPI.
Markers closely linked to the soybean rust resistance locus in G. tomentella T1 PI441001 and PI499939 were converted to TaqMan assays (assay ID 3289 and 3316) (Table S2). Both assays were used to determine the presence of the markers and copy number [28,29,31]. Putative F1 plants were evaluated for the presence of markers and their resistance to soybean rust using a detached leaf assay [32,33]. One soybean rust field population collected in North Carolina, RTP1, was used. A total of six fully expanded leaves were sampled following the procedure as described by Hao et al. (2024) [33]. Briefly, rust spores were collected using a Cyclone surface sampler (Burkard Manufacturing Co. Ltd., Rickmansworth, UK) and dissolved in 0.01% Tween 20 for approximately one hour before spraying directly onto the adaxial side of the leaves. Following rust inoculation, leaf samples were incubated for approximately two weeks in a growth chamber at 22 °C and 12-h light cycle before assigning a rust resistance rating. The rating includes lesion color (reddish-brown or tan-colored lesions) and lesion severity based on density and size from small visible lesion and limited number of lesions per leaf to large size lesions or high-density lesions per leaf [4,5].
We first selected five Syngenta soybean lines to evaluate their success rates across hybridization attempts, embryo rescue, and shoot regeneration. Each was crossed with two G. tomentella T1 accessions, PI441001 and PI583970, and one T2 accession, PI446958. Using PI441001 as the paternal progenitor, a range of 823 to 929 crossing attempts were conducted across all five soybean lines with three lines showing success rates above 49% in F1 embryo development (Table 2). When PI446958 was used as the paternal progenitor, four lines showed more than 50% crossing success rates with a range of 196 to 298 crossing attempts (Table 2). Using PI583970 as the paternal progenitor, all combinations reached 57% or higher success rates (Table 2). Across all combinations, approximately two embryos could be isolated per pod from the first wide crosses.
We also conducted embryo rescue (ER) using the five soybean lines as the maternal progenitor for crosses with G. tomentella T1 PI441001. No colchicine treatment was applied so that the rate of ER in tissue culture could be estimated. We found that SJ0225 and CC0653 showed higher rates of rescue success compared to the other three lines (Table 3). While HI2458 and HI0211 also showed high rates of crossing success and ER, they were susceptible to greenhouse pests, making them operationally challenging. Therefore, only SJ0225 and CC0653 were used to generate tetraploid soybean lines and used in the following wide crosses. Since no colchicine treatment was used to double the F1 genome, recovered F1 shoots were all sterile as expected.
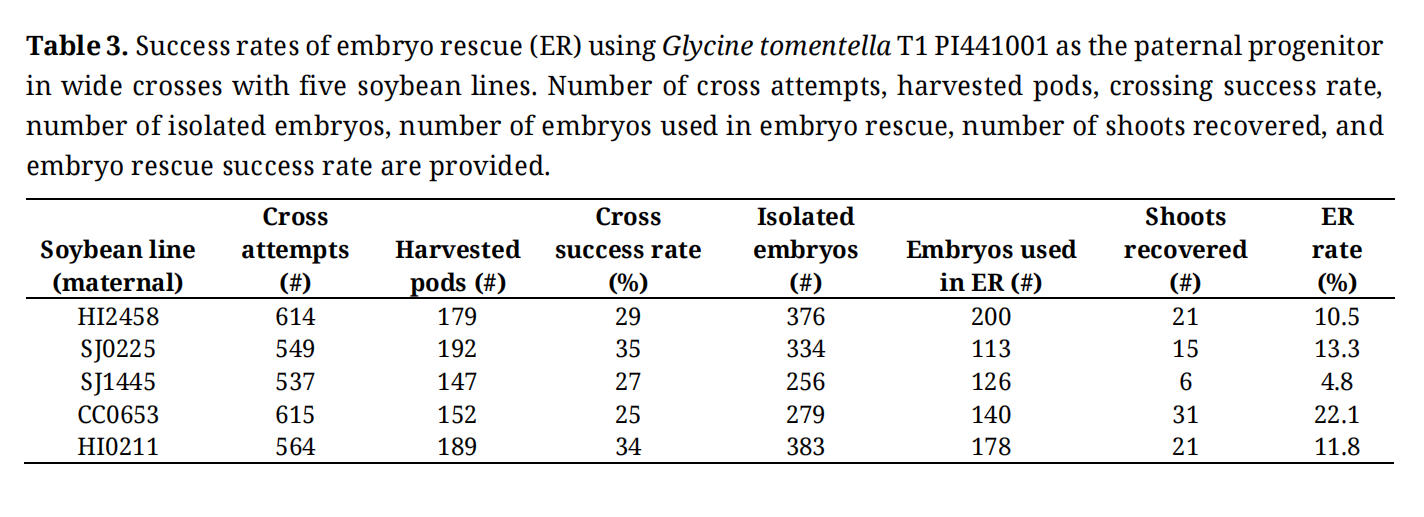 Table 3. Success rates of embryo rescue (ER) using Glycine tomentella T1 PI441001 as the paternal progenitor in wide crosses with five soybean lines. Number of cross attempts, harvested pods, crossing success rate, number of isolated embryos, number of embryos used in embryo rescue, number of shoots recovered, and embryo rescue success rate are provided.
Table 3. Success rates of embryo rescue (ER) using Glycine tomentella T1 PI441001 as the paternal progenitor in wide crosses with five soybean lines. Number of cross attempts, harvested pods, crossing success rate, number of isolated embryos, number of embryos used in embryo rescue, number of shoots recovered, and embryo rescue success rate are provided.
Tetraploid soybean lines were obtained through colchicine treatment of immature embryos and seed stocks were generated from multiple generations of self-pollination as described above. To differentiate from their progenitors, SJ0225 and CC0653, the two genome doubled soybean lines are referred to as SJ0225_T and CC0653_T. Although tetraploid soybean seeds are larger than their diploid progenitors, they are irregular in shape with visible dark marks (Figure 2A). When fully imbibed overnight, seed coats of SJ0225_T were darker than SJ0225 (Figure 2B). Germination rates of tetraploids were 65% compared to nearly 100% for diploids. Flow cytometry confirmed that the tetraploid stocks possessed doubled genomes (Figure 2C). Tetraploid soybean plants were also shorter. Based on our limited number of plants (n = 8) grown, we estimated that SJ0225_T or CC0653_T are approximately 20% shorter than their diploid progenitors. SJ0225_T and CC0653_T were also less fertile and produced fewer flowers with substantially less viable pollen, resulting in fewer self-pollinated seeds than their diploid progenitors (Figure 2D). Pollen from SJ0225_T and CC0653_T and their diploid progenitors were stained for viability. Four independent experiments were conducted and tetraploid lines were found to have an average of 22% of pollen stained, while nearly 100% of pollen was stained in the diploid progenitors. In our repeated experiments under greenhouse conditions, approximately 20 to 50 seeds could be harvested from a plant sown from tetraploid soybean lines, while approximately 300 seeds are readily harvested from their diploid progenitors.
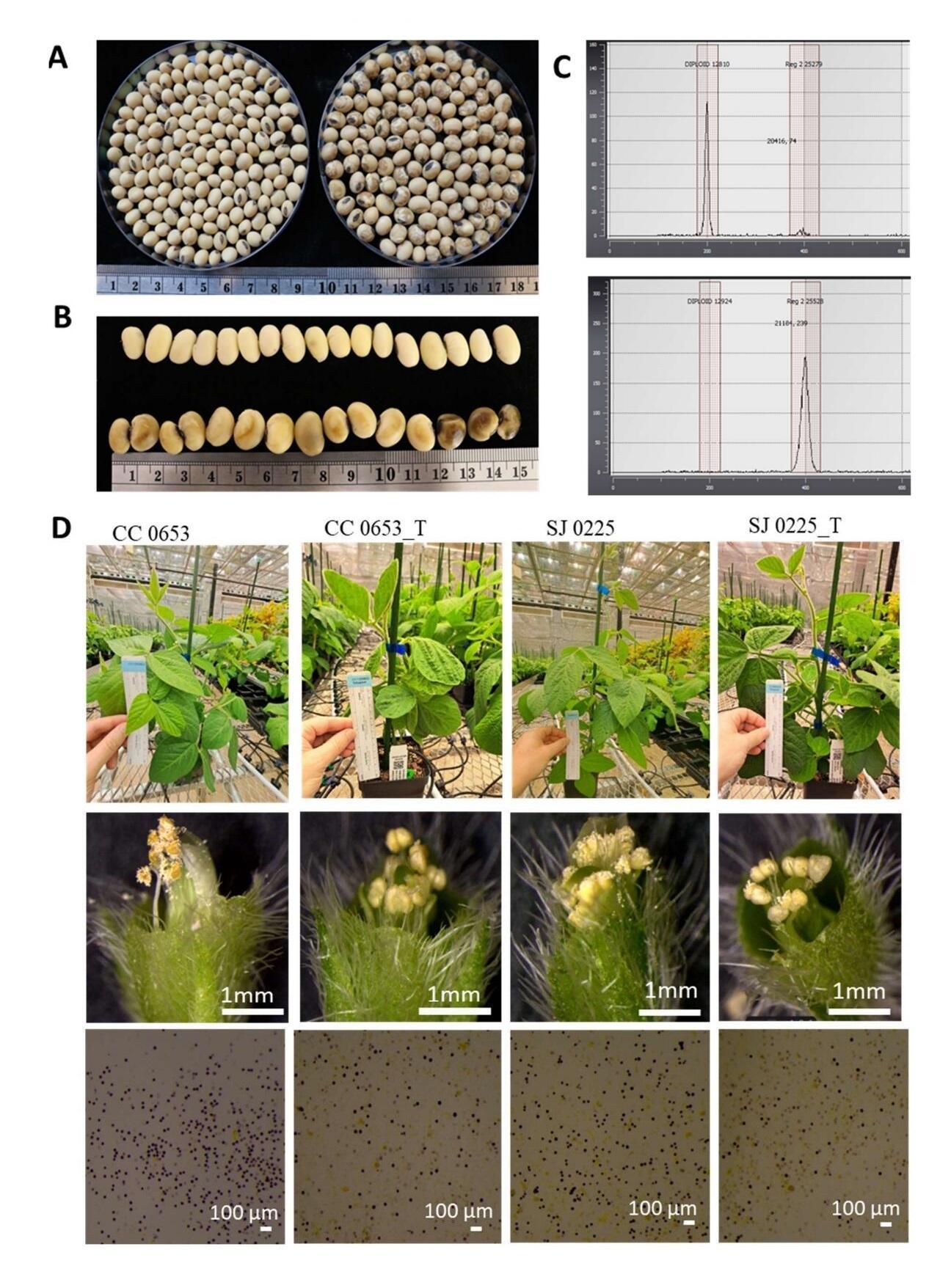 Figure 2. Comparison between soybean lines and their matching tetraploid lines (A) Dry seeds of SJ0225 (left) and SJ0225_T (right). (B) Imbibed seeds of SJ0225 (top) and SJ0225_T (bottom). (C) Flow cytometric analysis of SJ0225 (top) and SJ0225_T (bottom). Histograms display fluorescence intensity, representing the relative nuclear DNA content for each line. (D) (Top row) Greenhouse growth phenotypes illustrating that tetraploid soybean lines SJ0225_T and CC0653_T are visibly shorter than their diploid progenitors. (Middle row) Dissected flowers showing reduced pollen shed in tetraploids. (Bottom row) MTT staining revealing a higher frequency of non-viable (unstained) pollen grains in tetraploid lines compared to diploids.
Figure 2. Comparison between soybean lines and their matching tetraploid lines (A) Dry seeds of SJ0225 (left) and SJ0225_T (right). (B) Imbibed seeds of SJ0225 (top) and SJ0225_T (bottom). (C) Flow cytometric analysis of SJ0225 (top) and SJ0225_T (bottom). Histograms display fluorescence intensity, representing the relative nuclear DNA content for each line. (D) (Top row) Greenhouse growth phenotypes illustrating that tetraploid soybean lines SJ0225_T and CC0653_T are visibly shorter than their diploid progenitors. (Middle row) Dissected flowers showing reduced pollen shed in tetraploids. (Bottom row) MTT staining revealing a higher frequency of non-viable (unstained) pollen grains in tetraploid lines compared to diploids.
We hypothesized that the application of exogenous auxin, which delays endosperm cellularization, would be needed in crosses where the female has the higher ploidy. Wide crosses using tetraploid soybean, SJ0225_T, as the maternal progenitor and G. tomentella T1 PI441001 as the paternal progenitor resulted in an average of five crosses per plant. Wide crosses using diploid soybean as the maternal progenitor averaged 20 or more crosses per plant. Pods from tetraploid soybean wide crosses were also smaller than pods from diploid soybean wide crosses. Embryos from tetraploid soybean crosses did not develop without dicamba treatment (Figure 3A), making embryo rescue infeasible. In contrast, the embryos can be readily rescued and hybrid shoots generated when diploid soybeans are used. Crosses of tetraploid soybean (maternal) and diploid soybean (paternal) parents were also attempted but failed, similar to the cross between tetraploid soybean with G. tomentella T1 PI441001. These results support the hypothesis that tetraploid soybeans have a higher EBN than PI441001, and auxin treatment could improve the rates of embryo development. One time treatment of two dicamba concentrations (5 and 7 mg/L) promoted pod development (Figure 3B). Then, we assessed nine different regimes of dicamba applications with varying concentrations and durations. Gibberellin (GA) and kinetin were included in eight regimes but not the control treatment. In all different dicamba regimes, embryos were extracted to ER medium (Figure 3B). Dicamba concentrations ranging from 5–10 mg/L applied two or more non-consecutive days after pollination showed the highest success rates in embryo development (Table 4). Three or more daily applications of dicamba showed higher rates in shoot regeneration than one or two daily applications.
Putative F1 shoots were recovered from tetraploid soybean crossed with perennial Glycine. Two tetraploid soybean lines showed differences in the rate of embryo development. SJ0225_T exhibited lower rates of embryo development and recovery than CC0653_T when paired with G. tomentella T1 PI441001. However, SJ0225_T showed a higher rate of embryo development and similar recovery rate when crossed with G. tomentella T1 PI499939 (Table 5). Notably, we recovered fifteen embryos from 46 pollination attempts between SJ0225_T crossed with G. tomentella T5 PI509501 and recovered 1 F1 shoot. Additionally, 4 and 2 F1 embryos were recovered from crossing G. tomentella D3 PI505267 to SJ0225_T and CC0653_T respectively and 1 F1 shoot was recovered (Table 5). Embryo rescue success rates ranged from 6.7% to 25% among all the tested perennial accessions (Table 5), except for G. tomentella T1 PI483224. Four embryos isolated from SJ0225_T × PI483224 did not produce any F1 shoot.
Recovered F1 shoots were transplanted to soil, and they morphologically were intermediate between soybean and perennial Glycine parents. We noticed that F1 shoots derived from Glycine tomentella T1 PI499939 exhibit a morphology that closely aligns with the G. tomentella parent (Figure 4A). The morphological characteristics of an F1 shoot originating from diploid G. tomentella D3 PI505267 showed a distinct bias toward the G. max parent (Figure 4B).
To further evaluate the F1 shoots, three distinct analytical methodologies were employed. First, two TaqMan markers linked to soybean rust resistance were used to confirm F1 hybrids. The Taqman assays were negative in soybean lines SJ0225, CC0653, SJ0225_T, and CC0653_T. All recovered F1 plants were positive for these assays. Specifically, both assays were shown as 1 copy in 26 hybrid shoots derived from G. tomentella T1 PI441001 and PI499939 as the paternal progenitor, indicating the heterozygous state of the soybean rust resistance locus (Table 6).
Next, we used fluorescence in situ hybridization (FISH) to verify hybrids from tetraploid soybean wide crossed with perennial Glycine accessions PI441001 and PI499939. We observed 79 DAPI-stained chromosomes, 40 of which exhibited green fluorescence signals, indicating their soybean origin. The remaining 39 chromosomes were negative for the green signal indicating their origin from the perennial Glycine parents (Figure 4C).
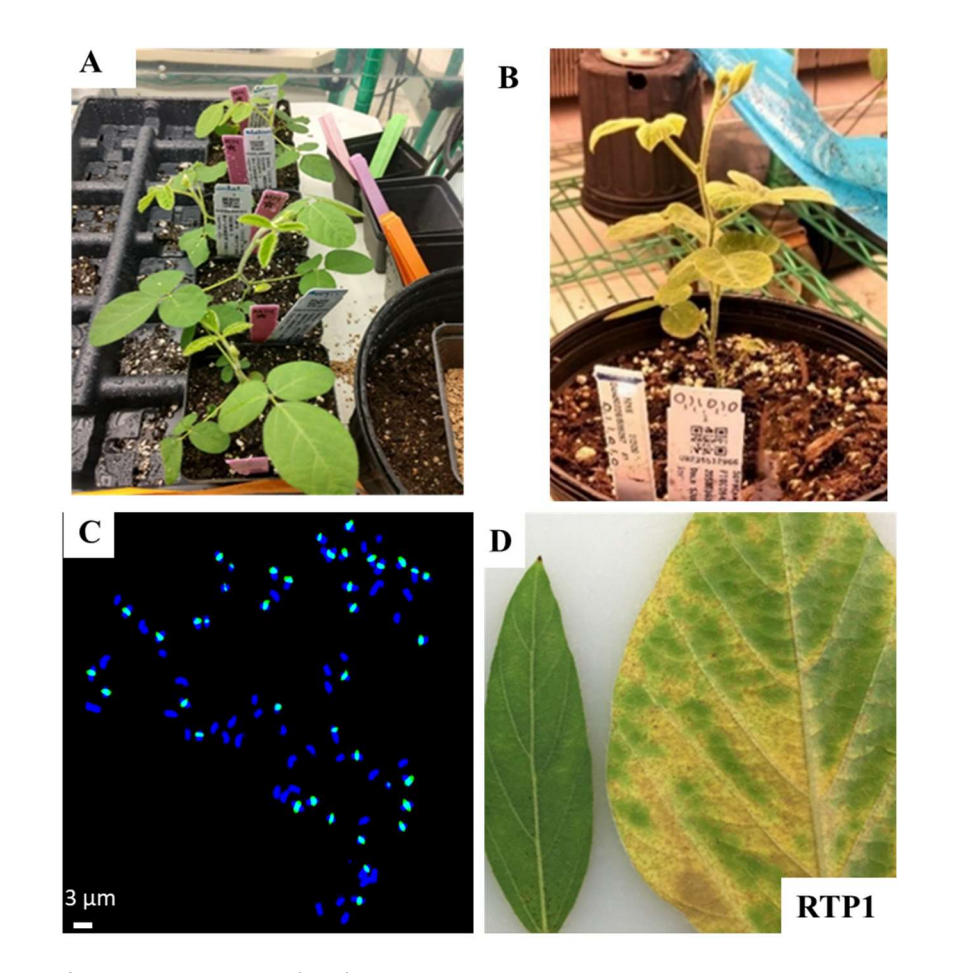 Figure 4. Characterizations of F1 plants from tetraploid soybean x perennial Glycine accessions. Greenhouse growth phenotypes of (A) F1 plants derived from SJ0225_T × PI499939 (T1); and (B) A single F1 plant derived from SJ0225_T × PI505267 (D3); (C) Chromosome spread of F1 from SJ0225_T × PI499939 (T1) detected by DAPI and Alexa Fluor® 488 stain; all chromosomes stained with DAPI are dark blue in color. Chromosomes with a light blue color were probed with a green-fluorochrome (Alexa Fluor® 488)-tagged with CentGm-1 and CentGm-2. Forty chromosomes were highlighted with green fluorescence, and 39 chromosomes were not labeled indicating their perennial Glycine origin. Scale bar is shown as labeled. (D) Soybean rust resistance observed on F1 plants inoculated with RTP1 rust spores showing small flecks without sporulation (left) in contrast to the soybean progenitor showing tan-colored lesions with profuse sporulation (right).
Figure 4. Characterizations of F1 plants from tetraploid soybean x perennial Glycine accessions. Greenhouse growth phenotypes of (A) F1 plants derived from SJ0225_T × PI499939 (T1); and (B) A single F1 plant derived from SJ0225_T × PI505267 (D3); (C) Chromosome spread of F1 from SJ0225_T × PI499939 (T1) detected by DAPI and Alexa Fluor® 488 stain; all chromosomes stained with DAPI are dark blue in color. Chromosomes with a light blue color were probed with a green-fluorochrome (Alexa Fluor® 488)-tagged with CentGm-1 and CentGm-2. Forty chromosomes were highlighted with green fluorescence, and 39 chromosomes were not labeled indicating their perennial Glycine origin. Scale bar is shown as labeled. (D) Soybean rust resistance observed on F1 plants inoculated with RTP1 rust spores showing small flecks without sporulation (left) in contrast to the soybean progenitor showing tan-colored lesions with profuse sporulation (right).
Finally, F1 hybrids were inoculated with soybean rust to evaluate the successful transfer of resistance. In leaf assays using isolate RTP1, F1 plants exhibited small, reddish-brown (RB) lesions with minimal to no sporulation. In contrast, the soybean parental lines displayed characteristic tan-colored (TAN) lesions with profuse sporulation, confirming that resistance from the perennial Glycine parent was successfully introgressed (Figure 4D; Table 6).
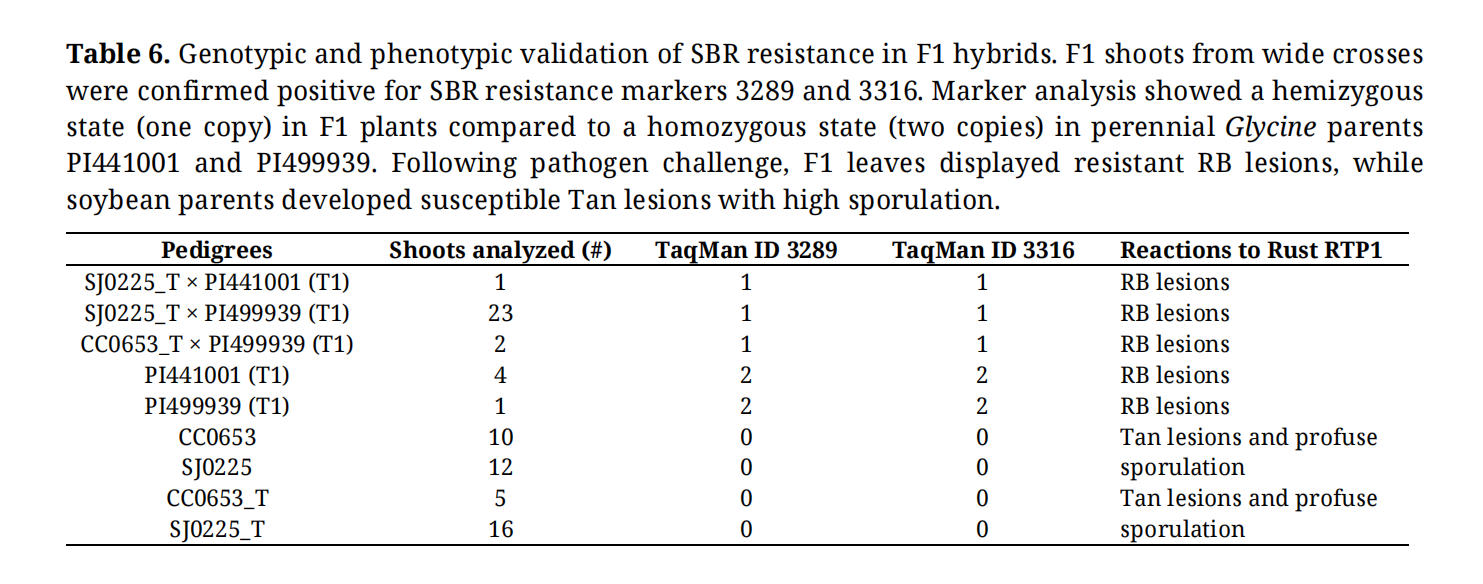 Table 6. Genotypic and phenotypic validation of SBR resistance in F1 hybrids. F1 shoots from wide crosses were confirmed positive for SBR resistance markers 3289 and 3316. Marker analysis showed a hemizygous state (one copy) in F1 plants compared to a homozygous state (two copies) in perennial Glycine parents PI441001 and PI499939. Following pathogen challenge, F1 leaves displayed resistant RB lesions, while soybean parents developed susceptible Tan lesions with high sporulation.
Table 6. Genotypic and phenotypic validation of SBR resistance in F1 hybrids. F1 shoots from wide crosses were confirmed positive for SBR resistance markers 3289 and 3316. Marker analysis showed a hemizygous state (one copy) in F1 plants compared to a homozygous state (two copies) in perennial Glycine parents PI441001 and PI499939. Following pathogen challenge, F1 leaves displayed resistant RB lesions, while soybean parents developed susceptible Tan lesions with high sporulation.
Our initial motivation to develop a tetraploid soybean method was due to the potential efficiency gained in time and number of crosses. Two G. tomentella accessions PI441001 and PI499939 were crossed to SJ0225_T and SJ0225 to directly measure efficiency of the tetraploid soybean method in the context of the modified Singh method. Using SJ0225_T as the maternal line, we conducted 32 wide crosses using PI441001 and recovered three F1 shoots after two and a half months, resulting in an efficiency of 9.4 F1 shoots per 100 wide crosses. For PI499939, 682 wide crosses were conducted, and twenty-five F1 shoots were recovered with a rate of 3.7 F1 shoots per 100 crosses. Using SJ0225 as maternal line, to generate F1 embryos for chromosome doubling, nearly 6,000 wide crosses were performed using accession PI441001, resulting in the recovery of seven doubled F1 (F1D) shoots. Due to the limited fertility of these F1D plants, an additional 1,800 backcrosses were conducted to produce seven BC1F1 shoots. The entire study encompassing the initial wide crosses, colchicine-mediated doubling, and subsequent backcrossing was completed in 461 days across two generations. When calculating efficiency by the number of GGDE shoots from the overall number of crosses conducted, the efficiency of the modified Singh method is 0.089 shoots per 100 crosses (e.g., seven shoots from 7800 wide crosses and backcrosses combined). For PI499939, despite the extensive scale of the hybridization efforts including over 3,000 wide crosses to obtain F1 embryos and 100 subsequent backcrosses, no BC1F1 shoots were successfully recovered. Overall, the tetraploid soybean method using SJ0225_T as maternal line achieved a range of 41-to-100-fold efficiency gain compared to the modified Singh method using SJ0225 as maternal line. The tetraploid soybean method also saved nearly 400 days to reach GGDE genome composition (Table 7).
When SJ0225_T was used as the female parent, ER was required to recover BC1F1 embryos to generate shoots. Embryo rescue was no longer needed to rescue BC2F1 embryos and BC2F1 mature seeds were harvested from pods (Figure 1B). In contrast, when SJ0225 was used as the female parent, ER was required to recover BC2F1 embryos to obtain BC2F1 shoots, while no ER was required to recover BC3F1 mature seeds (Figure 1A).
Using F1 shoots derived from SJ0225_T crossed with T1 PI441001 and PI499939 respectively, backcrosses using SJ0225 were conducted. BC2F1 seeds were obtained approximately 10 months after the first F1 cross. The BC2F1 seeds derived from PI441001 closely resembled G. max seeds in morphology but were substantially smaller in size than soybean (Figure 5A). BC2F1 seeds derived from PI499939 looked less like soybean and were smaller than the ones derived from PI441001 (Figure 5A). We genotyped 23 BC2F1 plants derived from PI499939 using soybean rust markers and found that 10 were positive for the markers, fitting the expected 1:1 ratio in a backcrossed population. We sampled root tips of three marker positive BC2F1 plants and conducted FISH to assess chromosome composition derived from SJ0225_T × PI441001 and SJ0225_T × PI499939. One individual showed two chromosomes missing the soybean centromere specific signal (Figure 5B), while two individuals showed one chromosome missing the signal (Figure 5C). We found all individuals sampled showed soybean rust specific markers co-localized with a soybean centromere specific marker suggesting integration of the soybean rust resistance locus in a soybean chromosome (insets of Figure 5B, 5C). These individuals were inoculated with soybean rust RTP1 and all developed RB lesions (Figure 5D, E). Notably, BC2F1 plants derived from PI441001 exhibited a highly resistant phenotype characterized by small, non-sporulating flecks. In contrast, BC2F1 progeny derived from PI499939 displayed a significantly higher lesion density and larger lesion sizes, suggesting a difference in the strength or mechanism of resistance provided by the two perennial accessions.
We further produced 51 BC4F1 plants derived from PI441001, of which 31 plants were positive for soybean rust resistance markers. The observed number of plants with the markers fits Mendelian segregation ratio of 1:1 as expected for a backcross population. One BC4F1 plant was selfed and produced 68 BC4F2 seeds. These 68 BC4F2 seeds were planted and genotyped. We found 14 individuals negative for soybean rust markers, 35 individuals with one copy of the markers indicating hemizygous state of the soybean rust locus, and 19 individuals with two copies of the markers for soybean rust resistance locus indicating homozygous state of the trait. This data fits the expected 1:2:1 Mendelian inheritance of the soybean rust resistance locus in a F2 population.
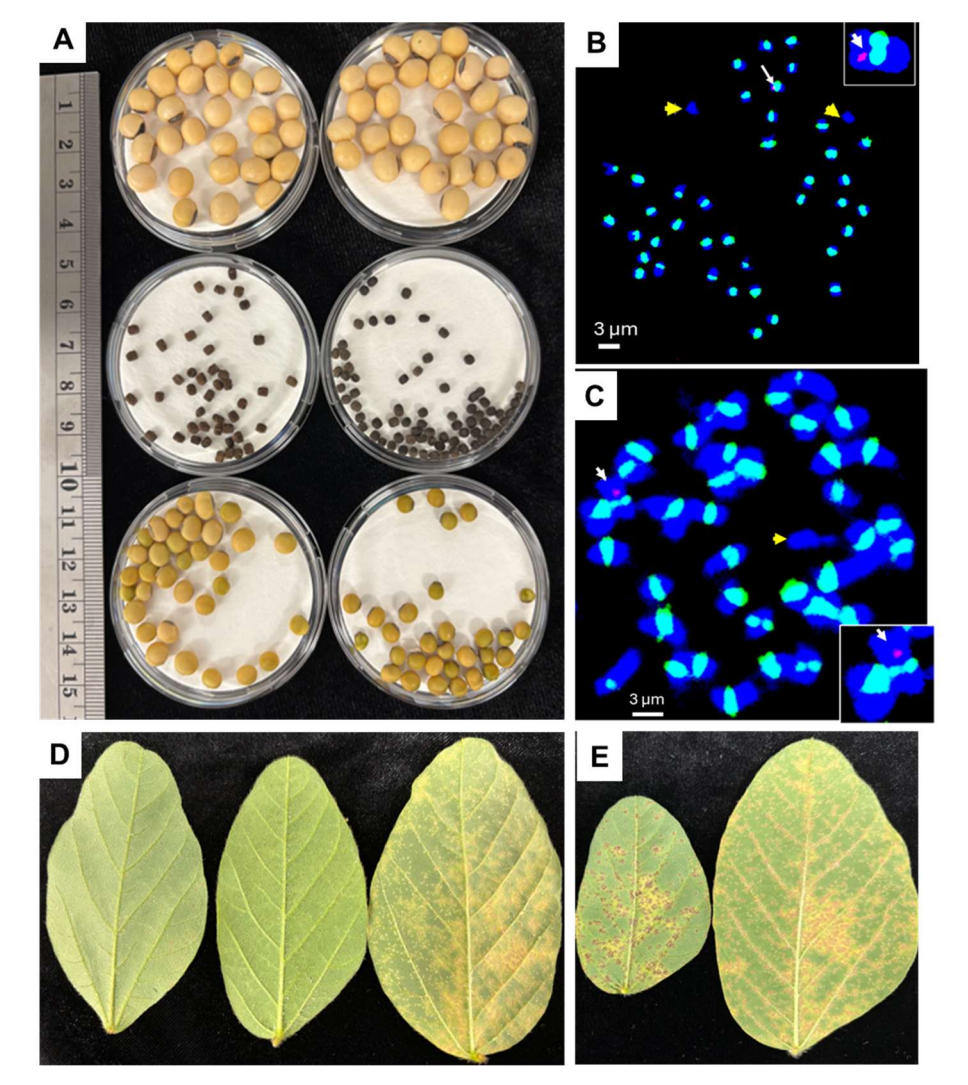 Figure 5. Characterizations of BC2F1 progeny: seed morphology, karyotypes, and soybean rust resistance. (A) Seed morphology comparison. Top panel: SJ0225; middle panel: perennial accessions PI441001 (left) and PI499939 (right); bottom panel: BC2F1 seeds derived from SJ0225_T × PI441001 (left) and SJ0225_T × PI499939 (right). Scale bar = 1 cm. (B) Cytogenetic analysis of BC2F1 progeny from SJ0225_T × PI441001. Chromosome spread subjected to fluorescence in situ hybridization (FISH). Green signals indicate soybean-specific centromeric probes (CentGm-1 and CentGm-2); red signals indicate a soybean rust (SBR)-specific marker. Yellow arrows denote chromosomes lacking soybean centromere signals (alien chromosomes). The white arrow and inset (top right) highlight the chromosome carrying the SBR-specific marker. (C) Cytogenetic analysis of BC2F1 from SJ0225_T × PI499939. Representative chromosome spread showing a single alien chromosome (unlabeled by green CentGm probes, yellow arrow). The white arrow and inset (bottom right) identify the chromosome carrying the red SBR-specific marker. (D) Phenotypic screening for SBR resistance of BC2F1 from SJ0225_T × PI441001. Leaves (left) exhibit resistant reddish-brown (RB) fleck lesions with no sporulation, whereas the SJ0225 parent (right) displays susceptible TAN lesions with profuse sporulation. (E) Phenotypic screening for SBR resistance of BC2F1 from SJ0225_T × PI499939. Leaf (left) showing macroscopic lesions with minimal sporulation, compared to susceptible TAN lesions with profuse sporulation in SJ0225 (right)./font>
Figure 5. Characterizations of BC2F1 progeny: seed morphology, karyotypes, and soybean rust resistance. (A) Seed morphology comparison. Top panel: SJ0225; middle panel: perennial accessions PI441001 (left) and PI499939 (right); bottom panel: BC2F1 seeds derived from SJ0225_T × PI441001 (left) and SJ0225_T × PI499939 (right). Scale bar = 1 cm. (B) Cytogenetic analysis of BC2F1 progeny from SJ0225_T × PI441001. Chromosome spread subjected to fluorescence in situ hybridization (FISH). Green signals indicate soybean-specific centromeric probes (CentGm-1 and CentGm-2); red signals indicate a soybean rust (SBR)-specific marker. Yellow arrows denote chromosomes lacking soybean centromere signals (alien chromosomes). The white arrow and inset (top right) highlight the chromosome carrying the SBR-specific marker. (C) Cytogenetic analysis of BC2F1 from SJ0225_T × PI499939. Representative chromosome spread showing a single alien chromosome (unlabeled by green CentGm probes, yellow arrow). The white arrow and inset (bottom right) identify the chromosome carrying the red SBR-specific marker. (D) Phenotypic screening for SBR resistance of BC2F1 from SJ0225_T × PI441001. Leaves (left) exhibit resistant reddish-brown (RB) fleck lesions with no sporulation, whereas the SJ0225 parent (right) displays susceptible TAN lesions with profuse sporulation. (E) Phenotypic screening for SBR resistance of BC2F1 from SJ0225_T × PI499939. Leaf (left) showing macroscopic lesions with minimal sporulation, compared to susceptible TAN lesions with profuse sporulation in SJ0225 (right)./font>
The genetic base of cultivated soybeans is restricted to annual Glycine max and G. soja, preventing breeders from utilizing much of the broader genetic diversity existing in perennial Glycine species [2,3]. Extremely low crossability and early pod abortion have limited the use of wild perennial species of the subgenus Glycine in soybean breeding programs [34]. Distinct gene pools for cultivated soybean exist based on varying success rates of hybridization among or between species: primary (GP-1), secondary (GP-2), tertiary (GP-3), and even further with quaternary (GP-4) [1–3]. GP-1 consists of Glycine max and its annual sister species, G. soja, which can cross with each other to produce fertile hybrids. Species in GP-2 of soybean have been postulated with no empirical evidence [2,3]. Although previously proposed that GP-3 of soybean consists of three species G. argyrea, G. canescens, and G. tomentella; only G. tomentella with the T1 (DDEE) genome type have progressed beyond amphiploid stages with exceptional expertise and combinations of hormone applications and culture media formulations [3,8,15–18,34–36]. All other perennial Glycine species and non-T1 G. tomentella lines belong to soybean GP-4 in that there have been no successful generation of viable hybrid plants, or if successful, did not progress to the amphiploid stage [3].
The most rate limiting step in previously reported successful intersubgeneric crosses between cultivated soybean and perennial wild relatives is the colchicine treatment. This step is crucial for the recovery of genome doubled F1s with extremely low fertility. Using the modified Singh method and two G. tomentella T1 accessions, our experiments showed a 0.089% success rate from wide cross to recover low fertility BC1F1s composed of the GGDE genome. To improve the overall rate of recovery of GGDE progeny, we developed a tetraploid soybean strategy enabled by dicamba treatment. Consistent with what has previously been reported, tetraploid soybean is difficult to work with due to many negative phenotypes, making it unsuitable in most wide cross breeding programs [26,27]. However, skipping the colchicine step in wide crosses and reducing experimental time are extremely valuable to commercial breeders. In this study, we successfully utilized tetraploid soybean enabled by dicamba treatment in the first wide crosses. We recovered three and twenty-five F1 plants with GGDE genome composition from G. tomentella T1 PI441001 and PI499939, respectively. Because genome composition of these F1 plants is equivalent to the BC1F1 plants in the modified Singh method, one life cycle, equivalent to more than one year, was saved.
We intend to introgress agriculturally important traits, such as soybean rust resistance, from perennial Glycine species into G. max. Such resistance could be stably inherited to advanced backcrossed generations. Based on FISH results, at least one plant at the BC2F1 generation derived from T1 PI441001 and PI499939 respectively appeared to have integrated soybean rust resistance locus onto soybean chromosomes. Our limited genetic analysis of the SBR trait showed the expected 1:1 ratio and 1:2:1 ratio for populations derived from backcrossing and selfing, respectively. Hence, we demonstrated that tetraploid soybean can be used to broaden the soybean genetic base and introgress agriculturally useful traits into soybean with increased efficiency.
Perennial G. tomentella non-T1 genome type accessions PI505267 and PI5059501 belong to the D3 and T5 genome groups, respectively. We did not find any previous reports on obtaining BC1F1 derived from D3 or T5 genome groups as measured by their genome composition, though hybrids were reported [3,34–37]. Based on the gene pool definitions, they belong to GP-4. F1 plants generated in this study using the tetraploid soybean method represent substantial progress from previous reports on intersubgeneric hybridization between G. max and perennial Glycine species. Specifically, these F1s are equivalent to BC1F1 of the modified Singh method. These results suggest that tetraploid soybeans enable broader gene flow between G. max and its perennial wild relatives, i.e., the tetraploid soybean expands crossability of G. max. These data also demonstrate the remarkable plasticity of the soybean genome and that the gene pool barrier can be overcome with the deployment of new technologies. We did not attempt to cross other perennial Glycine species to tetraploid soybean. However, it is plausible to hypothesize that more perennial Glycine species could be crossed to G. max using the tetraploid soybean method described in this study.
Global soybean production has shown a consistent upward trend (171 mt in 2001 to 396 mt in 2023–2024) (www.soystats.com). However, these gains will be difficult to maintain in the future and will likely not keep pace with the growing agricultural demand [2]. Numerous challenges call for a need to exploit the existing genetic diversity to broaden the genetic base of this important crop to consistently improve genetic gains, resilience to climate change, and resistance against diseases and pests [2,18,38]. With this tetraploid soybean method, we successfully broadened the gene pools of soybean beyond G. tomentella T1, making genetic diversity as well as many beneficial traits accessible for soybean improvement. Soybean diploid progenitors used were able to produce 300 seeds on average. However, at BC4F1 generation, only 68 seeds were harvested from one BC4F1 plant indicating severe linkage drag. The possibility remains that undesirable genes from wild parents may have been transferred together with agriculturally important traits, i.e., soybean rust resistance. Insufficient chromosome pairing between chromosome segments of soybean and its wild relatives can limit recombination frequency. These challenges need to be addressed to fully realize the value of this tetraploid soybean method. With more advanced genomic tools, agriculturally important traits such as disease resistance can be mapped and cloned in diverse perennial Glycine species. We can expect that employment of new breeding tools, such as genome editing and cisgenesis, will enable utilization of these useful traits from perennial relatives in cultivated soybean [39–42].
The following supplementary materials are available online, Table S1: Primers used to amplify soybean centromere-specific probes, Table S2: Primers and probes used in TaqMan assays.
All data generated from the study are available in the manuscript or supplementary files. The dataset of the study is available from the authors upon request.
Conceptualization: QL, CS, QQ; Methodology: QL, CS, QQ, TK, YW; Investigation and visualization: all authors; Writing: QL and CS; Review and editing: all authors.
All authors are employees of Syngenta Crop Protection LLC at the time of the study.
A patent application covering the information in this manuscript was submitted on 8/12/2020 and has been published as WO2022/035648.
We first thank S. Johnson and M. Hill for the tissue culture and crossing support, and C. Fan, J. McCuiston, W. Gu, and R. Carlin for TaqMan assays. We thank C. Baxter, L. Shi, M. Lee, R. Quadt, I. Jepson, and G. Wu for their leadership support, and B. Vernooij and E. Levy for project support. We acknowledge L. Mumm, J. Ho, S. Miles, K. Jones, K. Williamson, C. Leming and S. Kanjilal for licensing and guidance on intellectual property. We thank Beth Dalsing and Savithri Purayannur for their reviews. We thank David Walker, Jeffery Doyle, and Jacob Landis for insightful discussion and USDA-GRIN for sharing perennial Glycine accessions. We are also grateful to J.B. Landis and A. Liu for reviewing and editing this manuscript.
1.
2.
3.
4.
5.
6.
7.
8.
9.
10.
11.
12.
13.
14.
15.
16.
17.
18.
19.
20.
21.
22.
23.
24.
25.
26.
27.
28.
29.
30.
31.
32.
33.
34.
35.
36.
37.
38.
39.
40.
41.
42.
Schuller C, Dawson J, Wu Y, Madden A, Su X, Jiang Y, et al. Expanding the Genetic Base of Glycine max by Using Soybean Tetraploids in Intersubgeneric Crosses with Perennial Wild Relatives. Crop Breed Genet Genom. 2026;8(2):e260014. https://doi.org/10.20900/cbgg20260014.

Copyright © Hapres Co., Ltd. Privacy Policy | Terms and Conditions