Location: Home >> Detail
TOTAL VIEWS
Crop Breed Genet Genom. 2026;8(2):e260010. https://doi.org/10.20900/cbgg20260010
1 Plant Virus Department, Leibniz Institute DSMZ-German Collection of Microorganisms and Cell Cultures, 39108 Braunschweig, Germany
2 Plant Biotechnology and Bioinformatics, Institute for Cellular and Molecular Botany (IZMB), University of Bonn, 53115 Bonn, Germany
* Correspondence: Samar Sheat
Cassava brown streak disease (CBSD), caused by two distinct viruses, the cassava brown streak virus (CBSV) and the Ugandan cassava brown streak virus (UCBSV), leads to a destruction of tuberous roots from necrosis. Earlier research has identified cassava lines that are immune to the viruses and other cassava lines expressing resistance by restricting the viruses to the roots. To resolve early host responses associated with resistance, we performed a time-course RNA-seq analysis of CBSV graft-infected cassava (1, 5 and 10 days after inoculation; DAI) with two South American cassava genotypes showing contrasting resistance phenotypes; DSC 167, a highly resistant line not permitting virus replication, DSC 260, expressing resistance by restricting the virus infection to the roots and, a susceptible genotype (TMS 96/0304), conducive to systemic virus infection. Differential expression analyses of stems, roots, young and old leaves revealed a strong genotype- and tissue-dependent timing of transcriptional responses. Resistant genotypes mounted their largest response at 1 DAI, most prominently in stem tissue near the graft insertion (DSC 167: 436 DEGs; DSC 260: 335 DEGs), whereas the response of the susceptible TMS 96/0304 was observed only at 10 DAI (737 DEGs) and broadly characterized by a downregulation of defense-associated genes in roots. Enrichment and functional annotation linked early resistance responses to pattern-triggered immunity and stress signaling, including receptor-like kinases, calcium signaling (CaM/CML), hormone-associated regulators (ethylene, jasmonate and auxin/IAA), protein folding/chaperone networks (HSP90/HSP70/HSP40/DnaJ), and cell wall- and phenylpropanoid-related genes. Taken together, the results of this study indicate that a rapid, localized activation of immune signaling in immune/resistant cassava lines, is a hallmark of CBSV resistance. This instant response takes place in tissues adjacent to the entry point (stems) and precedes the systemic spread of the virus. The pathways and genes activated comprise pattern triggered immunity and stress signaling.
DAI, days after inoculation; DEG, differentially expressed genes; CBSD, Cassava brown streak disease; CBSV, cassava brown streak virus; CMD, cassava mosaic disease; Els, expression levels; PCA, principal component analysis; DEGs, differentially expressed genes
Cassava (Manihot esculenta Crantz) is a woody shrub from the family Euphorbiaceae and is cultivated in South America, Africa, and Asia [1,2]. It is a crucial food security crop in Africa, providing the nutritional foundation for millions [1,3]. In Africa, cassava is threatened by several diseases, and the cassava brown streak disease (CBSD) is having the most severe impact. Two distinct virus species cause CBSD; the cassava brown streak virus (CBSV) and the Ugandan cassava brown streak virus (UCBSV). They belong to the family Potyviridae, genus Ipomovirus, [4,5] have distinct single-stranded RNA genomes but induce similar symptoms in infected cassava [4]. Brown lesions on the stem, vein clearing and chlorosis symptoms, mostly on old leaves [6] and dark brown, necrotic areas in the tuberous roots, often throughout the entire tuber, are prominent signs of the disease [7,8]. This root necrosis can lead to a complete loss of the crop, presenting a significant threat to cassava production and harvest [9,10]. The viruses are transmitted by whiteflies (Bemisia tabaci) in a semi-persistent manner but the disease mainly spreads through the exchange of infected stem cuttings used for vegetative propagation [10,11].
Extensive efforts to develop CBSD resistant cassava lines only reached limited success because resistant varieties among African cassava were not available. Novel sources of virus resistance thus were identified in South American germplasm accessions [12] and the most promising lines were subsequently incorporated into breeding programs to create cassava with resistance against viruses causing CBSD and against viruses causing cassava mosaic disease (CMD) which is endemic wherever cassava is grown on the continent [12,13]. Highly stringent methods were used to screen for strong and durable resistance [12] and resistance phenotypes were identified belonging to two categories: highly resistant plants, represented by DSC 167, free of symptoms and with no virus replication or establishment, and resistant plants represented by DSC 260, with no aboveground symptoms and no detectable virus but virus invasions and symptoms confined to roots. Further studies [14] showed that the resistance also is associated with a restriction of the virus to the companion cells of the phloem. Interestingly, while there was no virus replication in DSC 167 from a scion growing on a DSC 167 rootstock providing the source of CBSV, the virus was translocated through the plant in the external phloem only while in a similar experiment with DSC 260, the external and internal phloem was invaded [14]. Up until today, a mechanistic explanation for this phenomenon is still pending.
RNA-seq is method of choice for analyses of molecular processes regulating gene expression. Comparative studies of resistant and susceptible cassava plants were done by RNA-seq [15] to analyze virus infections and cassava resistance response 28 days after introducing a mixed infection of CBSV and UCBSV. In a study by [16], RNA-seq analysis revealed that varieties considered resistant had a high number of overexpressed genes among which NAC genes from hormone signaling and phenylpropanoid pathway were found most significant. Research by [17], reported transcriptome analyses at early time points, 6 h and 1, 2, 5, and 8 days after graft-infection. Already 2 and 5 days after infection, many plant defense genes were differentially expressed in a resistant genotype, including translation elongation factors (eiF), NBS-LRR, heat shock proteins, and pathogenesis-related proteins [17]. Transcriptome studies [15,17] further suggested that earlier time points after infection should be conducted.
Resistant lines described earlier were included in our resistance screening and we questioned the virus resistance character of the cassava lines used [12]. We then based our RNA-seq analysis on the new sources of resistance we identified in cassava and observed over many years for their response against the viruses causing CBSD. Two South American germplasm accessions, the highly resistant line DSC 167 that does not permit virus replication, the resistant line DSC 260, that restricted the virus to the roots and the susceptible cassava, TMS 96/0304, were studied in an RNA-seq analysis performed with plant samples taken at 1, 5, and 10 DAI. Using transcript abundances as a proxy, gene expression at the defined time points was analyzed to determine differential gene expression highlighting genes activated during an initial response against virus infection and genes involved in regulating viral replication and movement in early resistance responses of cassava against CBSV following the workflow shown in Figure 1.
The cassava plants used in this study were obtained from the cassava collection of the DSMZ Plant Virus Department at the Julius Kühn-Institute (JKI) in Braunschweig, also comprising South American cassava germplasm provided by CIAT (Centro Internacional de Agricultura Tropical), Cali, Colombia. Three cassava genotypes were selected, each exhibiting a distinct response to CBSV infection: susceptible TMS 96/0304, root-restricted DSC 260, and resistant DSC 167 [12,18]. The virus isolate, CBSV-Mo83 (DSMZ PV-0949, GenBank accession FN434436) was maintained in infected TMS 96/0304 cassava plants and used as a source of buds for graft-infection. Cassava plantlets were propagated in vitro, transferred to soil, hardened and subsequently grown in the greenhouse under controlled conditions with temperatures between 26 °C and 30 °C, 12 h light and 60% humidity.
Plant InfectionsThe cassava lines DSC 167, DSC 260 and TMS96/0304 were infected via grafting using two buds per plant, from CBSV infected source plants to increase inoculum and ascertain virus infection, similar to the protocol outlined by [19]. Inoculation of the virus was conducted by grafting two axillary buds from CBSV-infected TMS 96/0304 plants onto the lower stems of 3-month-old rootstocks of each test group. To account for transcriptional responses from wounding plants of each genotype were grafted with buds from healthy plants of the same genotype.
In addition to the 54 experimental plants, three plants from TMS 96/304, DSC 167 and DSC 260 were grafted and maintained for further observation.
Monitoring Graft Success, Virus Inoculation and InfectionGraft success was monitored visually (bud viability, callus formation at the graft union and following continued growth of the recipient plant). Only plants with an intact graft union were included in tissue sampling. To validate the effectiveness of grafting to introduce CBSV and infect the plants, CBSV graft-infected TMS 96/0304, DSC 167 and DSC 260 plants kept aside, were inspected for virus symptom development and tested by qRT-PCR. Furthermore, RNA-seq data were analyzed for virus reads by mapping to the CBSV-Mo83 genome sequence (FN434436.1).
Sample CollectionSamples of grafted plants were collected at three time points: 1, 5, and 10 days after inoculation (DAI). The earliest 1 DAI time point was selected to capture immediate host transcriptional responses at the primary graft interaction zone to also account for wound reactions.
Samples were taken from the stem below the bud insertion, the roots, the youngest fully expanded leaf (YL), and an older leaf from the lower stem (OL). For each genotype, time point, plant part, and treatment, nine plants were individually sampled (Table S1) and samples taken from stem, YL, OL and roots of 3 plants were pooled for RNA extraction. Thus, three biological replicates, defined as A, B, and C, for each genotype were collected and flash frozen in liquid nitrogen prior to RNA extraction.
RNA ExtractionFor virus detection, small sections of cassava tissue were transferred into 2 mL microcentrifuge tubes preloaded with stainless-steel beads, rapidly frozen in liquid nitrogen (N2) and kept at −80 °C until processing. The material was homogenized using a TissueLyser (QIAGEN, Germany). Total RNA was then isolated with an Epoch RNA extraction kit (Epoch Life Sciences, Missouri City, TX, US) essentially following the manufacturer’s instructions.
For RNA sequencing, frozen plant samples were ground in the TissueLyser (QIAGEN, Germany) and RNA was extracted using the RNeasy Plant Mini Kit (QIAGEN), following the manufacturer’s protocol. Residual DNA was removed through a DNase I treatment performed following a protocol provided by TAL (Transcriptome Analysis Laboratory, Göttingen, Germany). The reaction mixture utilized for the DNase I treatment included 40 µL of extracted RNA, 20 µL of 10× incubation buffer, 4 µL of DNase I (10 U/µL), 2 µL of RNase OUT (40 U/µL), and 134 µL of RNase-free water. The mixture was incubated at 37 °C for 20 min. Following, 100 µL of RNase-free water and 300 µL of phenol/chloroform were added. The samples were vortexed and centrifuged at 15,304× g for 2 min at room temperature. The upper phase containing RNA was transferred to a new 1.5 mL tube for RNA precipitation adding 30 µL 3 M sodium acetate (pH 4.8) and 300 µL isopropyl alcohol. Following vortexing and a brief centrifugation, the tubes were incubated at −20 °C for 30 min, then centrifuged for precipitation (15,304× g for 30 min at 4 °C) after which the supernatant was removed and the RNA pellet was washed twice with 1 mL of 75% ethanol. After the final wash, residual ethanol was carefully removed, and the pellet was air-dried and finally resuspended in 30 µL of RNase-free water. RNA was quantified using a NanodropTM2000 (PEQLAB, Erlangen, Germany) and by a checking the integrity in an 1% agarose gel. RNA samples were stored at −80 °C.
Quantitative Reverse Transcription Polymerase Chain Reaction (qRT-PCR)Detection of CBSV was carried out using a one-step TaqMan qRT-PCR assay with COX (cytochrome oxidase) included as an internal plant RNA quality/control. The protocol of Sheat et al. (2019) [13] was followed using a Maxima Probe/ROX qPCR Master Mix (Thermo Fisher), M-MLV RT, and CBSV- and COX-specific primers/probes. QRT-PCR was performed in a qTOWER3 (Analytik Jena) using 43 °C for 30 min for cDNA synthesis followed by an initial denaturation at 95 °C for 2 min to amplify for 40 cycles of 95 °C (15 s), 60 °C (30 s), and 72 °C (30 s). Results were reported as presence/absence based on Ct values, with COX confirming reaction performance and RNA integrity.
cDNA Library and RNA-seqThe cDNA libraries were built using the Illumina Stranded mRNA Prep Kit (Illumina, USA) following the manufacturers protocol Illumina [20]. From total RNA preparations, mRNA was purified using oligo(dT) magnetic beads and the enriched mRNA was then used to create the cDNA library for Illumina sequencing. The final cDNA library samples were evaluated for integrity and quality using the Qubit DNA High Sensitivity assay (Invitrogen, Thermo Fisher Scientific, Bremen, Germany) and the Agilent 2100 Bioanalyzer DNA 1000 High Sensitivity Kit (Agilent, CA, US).
Prior to sequencing, the cDNA libraries were diluted to 2 nM to ensure uniform concentrations. Sequencing was done on an Illumina NextSeq2000 using P3 and P2 FlowCells (paired-end 2 × 150 nt reads, 200 cycles).
Data AnalysisThe initial data analysis was conducted using Geneious Prime version 2025.0.2 (Dotmatics, Boston, MA, US) to import raw reads (FASTQ) after removing Illumina adapters and indexes using the R package BBDuk (version 38.84; [21]). Virus reads were identified by mapping against the CBSV-Mo83 genome sequence (FN434436.1, NCBI) aligning the non-used reads to the cassava reference genome sequence AM560-2, version 8 (GCF_001659605.2, NCBI), to cassava mitochondria (NC_045136.1, NCBI) and chloroplast (NC_010433.1, NCBI) sequences using the RNA mapper implemented in Geneious. Gene expression levels were calculated from paired-end reads mapped to the cassava reference genome. A CSV file, comprising transcripts per million (TPM) values for all 54 samples of one tissue type dataset, was imported into R to generate PCA plots using the R tool ggplot2 [22]. To explain potential outliers in the dataset, residual rRNA was checked for each sample using the Python script rRNA_check.py (version v0.1; [23]. DESeq2 [24] was used to determine the expression levels among infected and healthy samples of the same genotype and time point. Normalizing for sequencing depth and RNA composition was done by the median of ratios method which is implemented in DESeq2 [24,25]. The resulting CSV file was exported with columns for gene ID, protein ID, log2 fold change (FC), and adjusted p-value. Transcripts with an adjusted p-value > 0.05 and log2FC between −1.0 and 1.0 [26,27] were excluded to focus on major, significant differences. Bar charts were calculated to illustrate the total number of differentially expressed genes (DEGs). The protein IDs of DEGs were used in NCBI Batch Entrez [28] in which they got translated into proteins functions and subsequently presented in terms of appearance and regulation. Pathway analyses were conducted by translating the protein IDs of the cassava samples into the corresponding gene IDs of Arabidopsis thaliana using BLAST against the Arabidopsis thaliana proteome. KEGG pathway analyses were applied by using the gene IDs of Arabidopsis thaliana, generated from the biological DataBase network [29] to perform a KEGG enrichment analyses in R. Temporal expression pattern plots, using the log2FC values from the most conspicuously differentially expressed genes encoding protein candidates were generated. The changes in the log2FC values over time in stem, root, and leaf tissues were graphically represented in diagrams to compare the expression changes among the genotypes.
For additional analysis, a two-factor DESeq2 analysis was implemented to compare virus-infected genotypes, highly resistant, root-restricted and susceptible genotypes. The expression levels of healthy vs. infected genotypes were compared using stem and root reads at 1 and 10 DAI. DEGs encoding proteins IDs with an adjusted p-value > 0.05 and log2FC ratios between −1.5 and 1.5 were excluded, thereby focusing exclusively on significant up- or downregulated proteins. The number of DEGs proteins was counted and protein functions were determined via NCBI Batch Entrez. The two-factor DESeq2 analysis was compared with the one-factor DESeq2 analysis to evaluate whether similar DEGs proteins were found. Moreover, a three-factor analysis using DESeq2 together with a Likelihood Ratio Test (LRT) was implemented to directly compare the three genotypes among each other. In a first DESeq2 analysis, infected replicates were compared to healthy replicates of the same genotype. In the subsequent LRT step, the three genotypes were compared among each other. The susceptible TMS 96/0304 genotype was taken as the basis. During the analysis the adjusted p-value threshold of <0.05 was directly implemented to only filter significant differentially expressed genes. After the analysis, a CSV file was generated, containing protein ID, adjusted p-value, and log2FC. The number of proteins was counted, and protein functions were determined via NCBI Batch Entrez. The results of all DESeq2 calculations were compared and analyzed.
To study early transcriptome responses of cassava to infection with CBSV, graft inoculations were conducted and a solid graft union between virus source (2 buds from CBSV infected source plants) and rootstock ensured virus transfer [13,16].
Graft-inoculation to infect DSC 167 and DSC 260 with CBSV failed to induce symptoms on both lines indicating for their resistance status. In contrast, CBSV infection of the susceptible cassava TME 96/0304 became visible on newly emerging leaves at approximately 10 DAI and all three TME 96/0304 plants kept for virus observation developed typical CBSV symptoms approximately 1 month after infection (Figure 2B,C) while DSC 260 and DSC 167 plants remained symptom free (Figure 2D–G).
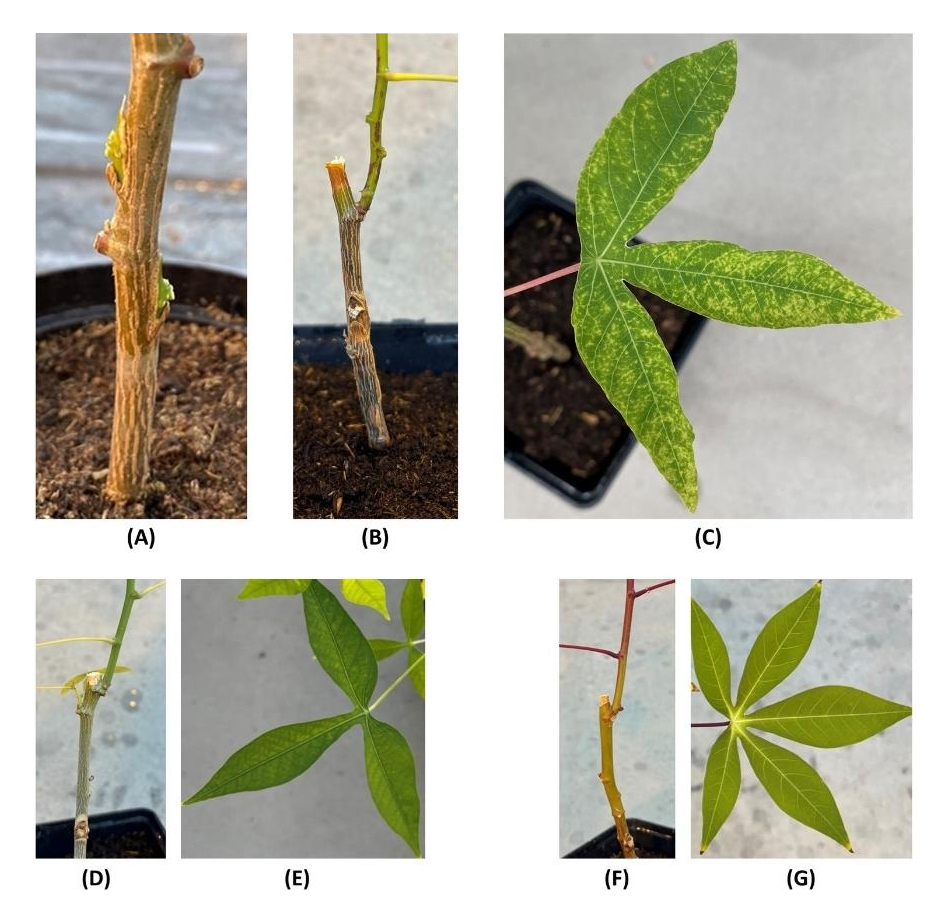 Figure 2. CBSV-infection by bud-grafting. (A,D,F) graft union on the test plants; (B,C) symptom expression of TMS 96/0304 after graft-infection with CBSV-Mo83 approximately 1 month after inoculation. Brown lesions (B) alongside the stem and chlorosis and mosaic symptoms on leaves (C) are typical signs of CBSD. Graft union in DSC 167 (D) and DSC 260 (F) confirms successful grafting/inoculation. Absence of leaf symptoms in DSC 167 (E) and DSC 260 (G) indicates virus resistance.
Figure 2. CBSV-infection by bud-grafting. (A,D,F) graft union on the test plants; (B,C) symptom expression of TMS 96/0304 after graft-infection with CBSV-Mo83 approximately 1 month after inoculation. Brown lesions (B) alongside the stem and chlorosis and mosaic symptoms on leaves (C) are typical signs of CBSD. Graft union in DSC 167 (D) and DSC 260 (F) confirms successful grafting/inoculation. Absence of leaf symptoms in DSC 167 (E) and DSC 260 (G) indicates virus resistance.
qRT-PCR confirmed presence of CBSV at 10 DAI in TMS 96/0304, albeit close to the detection threshold, when CBSV was detected in two pooled root samples indicating early viral accumulation in root tissues. Viral RNA was not detected in any of the samples from DSC 167 and DSC 260. All three TME 96/0304 plants left for long-term observation tested positive for CBSV while DSC 167 and DSC 260 remained negative (Table S2).
Mapping RNA-seq Reads Against the Cassava Brown Streak Virus Genome SequencePost sequencing analysis revealed good quality and quantity of the clean reads throughout all samples.
At 10 DAI, CBSV reads were detected in stem and root samples of TMS 96/0304, with the highest read counts in roots (Table 1, approximately 0.056% and 0.052% of total reads, respectively). In stem tissue, CBSV reads were detected in two of three replicates (0.0089% and 0.0025% of total reads). Despite the low number of virus-reads, qRT-PCR revealed CBSV-Mo83 in root replicates A and B of TMS 96/0304 (Table S2). There were no CBSV reads detected in DSC 260. RNA-seq of the highly resistant genotype DSC 167, revealed CBSV reads in one library (Stem A: 78,492 reads; approximately 0.11% of total reads; Roots A: 1,192 reads; approximately 0.0031% of total reads) while all other RNA-seq reads were devoid of CBSV sequences (Table 1). Complete mapping metrics for all libraries are provided in (Tables S3–S6).
Virus infection is spatially heterogeneous during early CBSV establishment thus, viral read counts at 10 DAI were interpreted as evidence of presence/absence in the sampled tissue and not as a quantitative proxy of the plant infection status. This is particularly important for the virus-resistant DSC 167, which, despite viral RNA (Table 1) translocating through phloem tissues from virus source (graft) to sink does not support virus replication (Table S2).
Clustering of Samples by Tissue TypePrincipal component analysis (PCA) of TPM values classified each sample according to its specific tissue type (Figure 3) and assigned RNA-seq from stem, root, young leaf (YL), and old leaf (OL) tissues to their respective groups demonstrating strong coherence among biological replicates. Several outliers were identified (Figure 3, dotted circles) and in root samples this was likely due to low RNA quality (assessed by Qubit and Bioanalyzer). The outliers recorded for YL and stem samples were attributed to biological variability since similar rRNA content was recorded.
Additional PCA plots separate for each tissue type revealed a read distribution according to cassava genotype (FigureS1–S4). PCA plots of stem, root and YL tissue samples of the two resistant lines showed some overlap but were well discriminated from those of TMS 96/0304. Since the PCA plot of the OL samples displayed a markedly different pattern, reads could not be explicitly assigned to a particular cassava genotype.
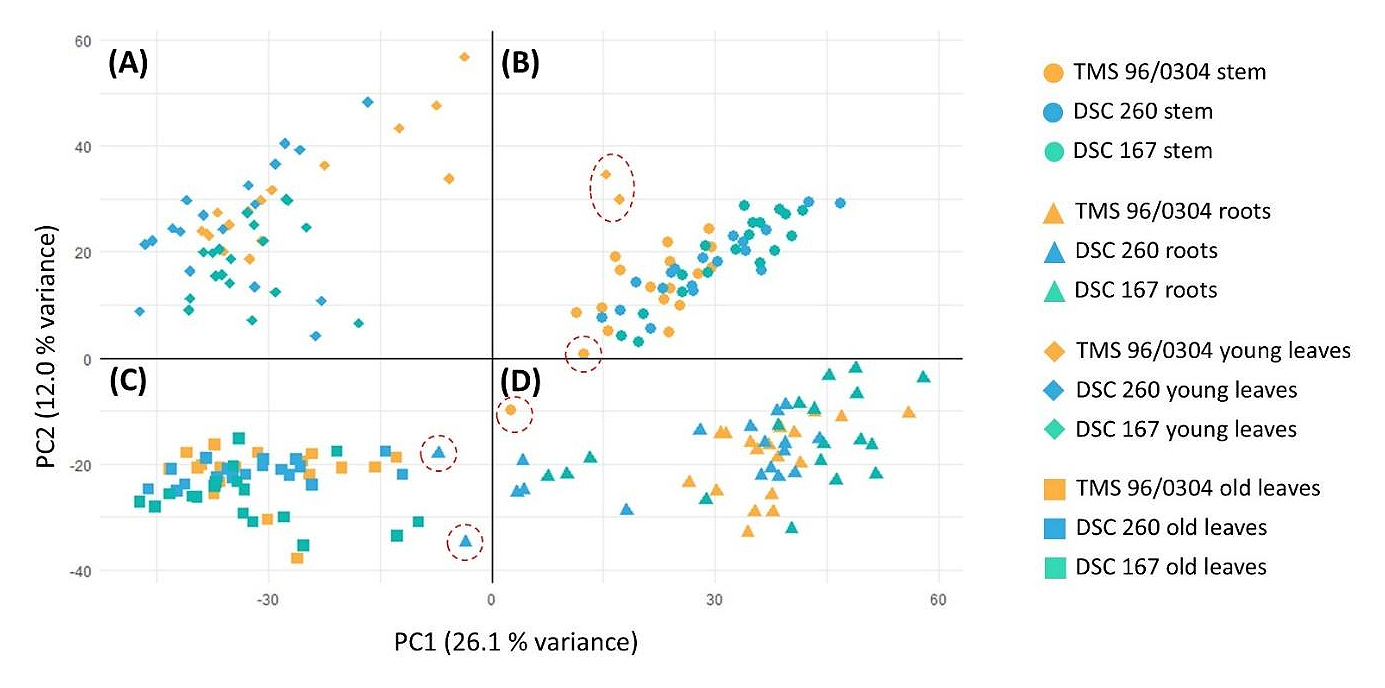 Figure 3. Principal component analysis (PCA) of all individual biological replicates. Replicates are grouped according to tissue type. (A) Replicates derived from young leaves cluster together. (B) Stem-derived replicates form a distinct cluster, with two young leaf samples appearing as outliers (orange diamonds: TMS 96/0304 healthy, replicate A at 1 DAI; TMS 96/0304 infected, replicate A at 1 DAI) and one stem outlier (orange circle: TMS 96/0304 infected, replicate A at 1 DAI). (C) Replicates from old leaves cluster together and include two root-derived outliers (blue triangles: DSC 260 healthy, replicate B at 5 DAI; DSC 260 healthy, replicate A at 10 DAI). (D) Root replicates form a separate cluster, including one stem-derived outlier (orange dot: TMS 96/0304 healthy, replicate A at 1 DAI). Dot, stem; triangle, roots; rhombus, young leaves; square, old leaves; orange, susceptible TMS 96/0304; blue, root restricted DSC 260; green, resistant DSC 167; outliers, red dotted circles; PC1 is 26.1 % variance; PC2 is 12.0 % variance. ● stem; ▲ roots; ◆ young leaves; ■ old leaves.
Figure 3. Principal component analysis (PCA) of all individual biological replicates. Replicates are grouped according to tissue type. (A) Replicates derived from young leaves cluster together. (B) Stem-derived replicates form a distinct cluster, with two young leaf samples appearing as outliers (orange diamonds: TMS 96/0304 healthy, replicate A at 1 DAI; TMS 96/0304 infected, replicate A at 1 DAI) and one stem outlier (orange circle: TMS 96/0304 infected, replicate A at 1 DAI). (C) Replicates from old leaves cluster together and include two root-derived outliers (blue triangles: DSC 260 healthy, replicate B at 5 DAI; DSC 260 healthy, replicate A at 10 DAI). (D) Root replicates form a separate cluster, including one stem-derived outlier (orange dot: TMS 96/0304 healthy, replicate A at 1 DAI). Dot, stem; triangle, roots; rhombus, young leaves; square, old leaves; orange, susceptible TMS 96/0304; blue, root restricted DSC 260; green, resistant DSC 167; outliers, red dotted circles; PC1 is 26.1 % variance; PC2 is 12.0 % variance. ● stem; ▲ roots; ◆ young leaves; ■ old leaves.
Upon graft infection, the resistant cassava genotypes (DSC 167, DSC 260) showed a rapid response to CBSV infection at 1 DAI which was characterized by a steep increase of gene expression in stem, roots, YL, and OL (Figure 4A). A large number of differentially expressed genes (DEGS) were found in YL tissue, 405 for DSC 260 and 135 for DSC 167 and in stem tissue, 335 for DSC 260 and 436 for DSC 167 (Table S7). At this time point, there was no striking response from the susceptible cassava TMS 96/0304 (Figure 4A) with only up to 47 DEGS in root and old leaf tissue. In contrast, at 10 DAI, the number of DEGs in TMS 96/0304 increased strongly (up to 1124 in OL (Table S7) while the number of DEGs in DSC 260, and DSC 167 were considerably low (Figure 4C) with lowest 9 to 0 DEGS (Table S7). Interestingly at time point 5 DAI, with the exception of OL in DSC 167, there were no considerable differences in gene expression profiles recorded (Figure 4B). Since the distribution of OL reads in the PCA plot and the number of DEGs were not informative, RNA-seq data from OL were exempted from further analysis.
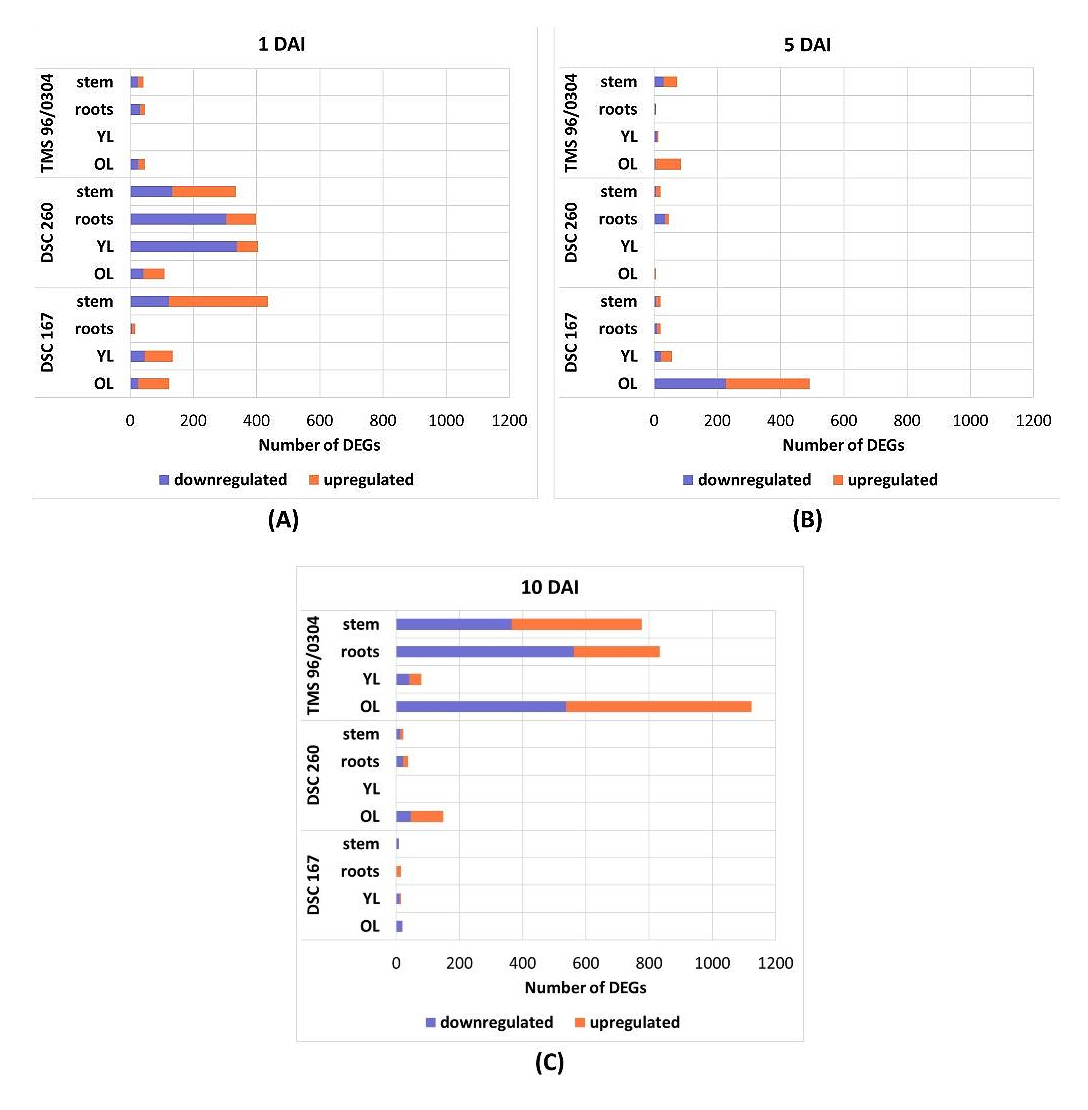 Figure 4. Bar charts of DEGs for each genotype and tissue. Thresholds for adjusted p-value < 0.05 and |log2 fold change| > 1. Upregulated genes (orange), downregulated genes (blue). (A) 1 DAI; a high number of DEGs in DSC 260 and DSC 167 plants. (B) 5 DAI; no significant DEG except in OL of the resistant genotype DSC 167. (C) 10 DAI; significant DEG in the susceptible genotype TMS 96/0304 while no changes recorded for the resistant cassava lines. DAI, days after inoculation; DEGs, differentially expressed genes; TMS 96/0304, susceptible; DSC 260, root-restricted; DSC 167, resistant.
Figure 4. Bar charts of DEGs for each genotype and tissue. Thresholds for adjusted p-value < 0.05 and |log2 fold change| > 1. Upregulated genes (orange), downregulated genes (blue). (A) 1 DAI; a high number of DEGs in DSC 260 and DSC 167 plants. (B) 5 DAI; no significant DEG except in OL of the resistant genotype DSC 167. (C) 10 DAI; significant DEG in the susceptible genotype TMS 96/0304 while no changes recorded for the resistant cassava lines. DAI, days after inoculation; DEGs, differentially expressed genes; TMS 96/0304, susceptible; DSC 260, root-restricted; DSC 167, resistant.
Protein IDs of the DEGs were mapped for functional annotation to entries in the NCBI Protein database [30]. The resulting genes were allocated to distinct gene clusters associated with hormonal regulation, transcriptional regulation, stress response, defense and immunity, cell structure, and signal transduction. A total of 24 gene categories were formed within these gene clusters, and the functions of the individual genes within each category were identified (Table S7).
For each time point (DAI), genotype and tissue, the DEGs belonging to a given gene category were counted. The resulting heatmap (Figure 5) illustrates the net trend within each group, calculated as the difference between the number of up- and downregulated genes. Their expression was weighed against each other, even if they partially leveled each other out. (e.g. −2 indicating two more downregulated than upregulated genes). This approach highlighted regulatory patterns rather than accounting for absolute expression levels and the magnitude of differential expression.
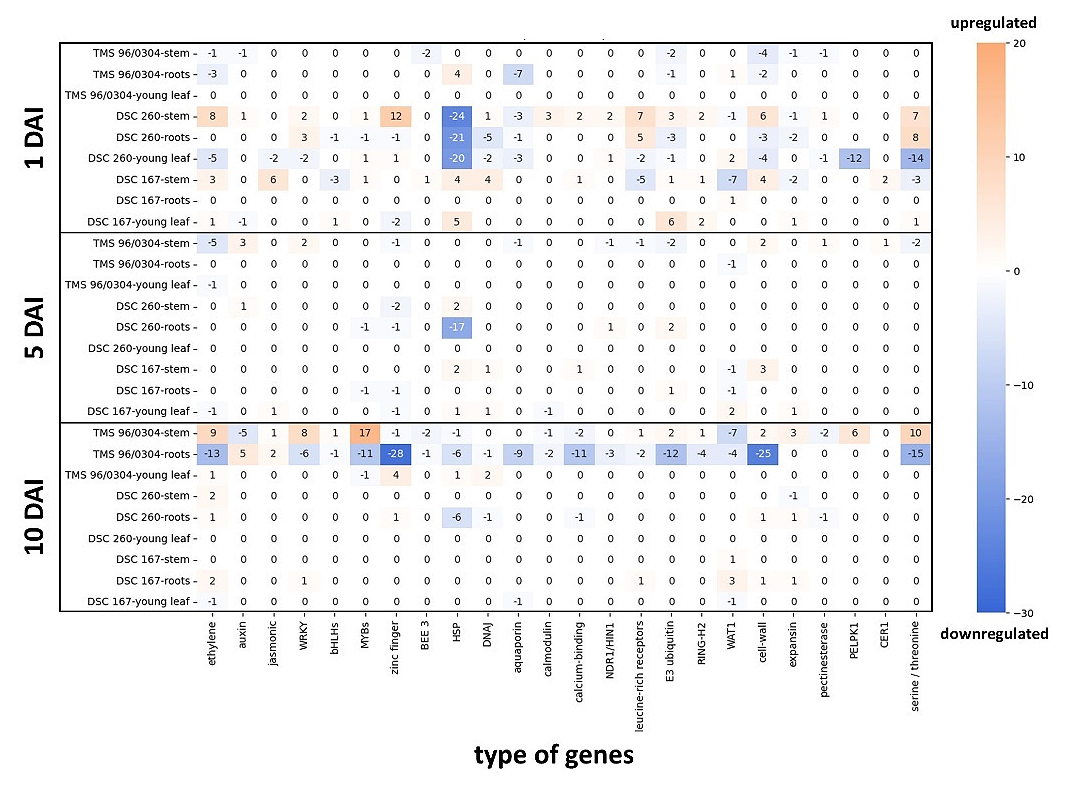 Figure 5. Heatmap of DEGs at time points (DAI), genotypes, and tissue types plotted against plant immune response genes. Threshold for adjusted p-value < 0.05 and |log2 fold change| > 1. Downregulation (blue), upregulation (orange). Strong upregulation in the stem tissue of the resistant genotypes DSC 260 and DSC 167 at 1 DAI Significant downregulation of almost all gene categories at 10 DAI in roots and upregulation in stems of the virus infected TMS 96/0304 at 10 DAI. DAI = days after inoculation; TMS 96/0304, susceptible; DSC 260, root-restricted; DSC 167, resistant.
Figure 5. Heatmap of DEGs at time points (DAI), genotypes, and tissue types plotted against plant immune response genes. Threshold for adjusted p-value < 0.05 and |log2 fold change| > 1. Downregulation (blue), upregulation (orange). Strong upregulation in the stem tissue of the resistant genotypes DSC 260 and DSC 167 at 1 DAI Significant downregulation of almost all gene categories at 10 DAI in roots and upregulation in stems of the virus infected TMS 96/0304 at 10 DAI. DAI = days after inoculation; TMS 96/0304, susceptible; DSC 260, root-restricted; DSC 167, resistant.
Upon CBSV infection, plant immune response genes showed notable alterations already at 1 DAI in the resistant cassava lines. In DSC 167 an increased gene expression of ethylene, jasmonic acid, HSPs, DnaJ and, cell wall biosynthesis was observed in stem tissues as well as an increase of HSP and E3 ubiquitin in leaves. In contrast, no significant differential expression was detected in root tissues at this time point. In contrast, DSC 260 showed an increased level of ethylene, zinc finger, calmodulin, leucine rich receptors, cell wall biosynthesis, and serine/ threonine kinases in stem tissue while a dramatic downregulation of HSP was recorded for stem, leaf and root tissues (Figure 5). At 1 DAI, there were no differential expression patterns recorded for TMS 96/0304 responding with most critical gene expression changes only at 10 DAI and most prominently in root tissues with almost all immune response-associated genes downregulated. In contrast, RNA-seq of stem tissues showed a notable upregulation of genes involved in ethylene biosynthesis, WRKY, MYB, cell-wall, pectin esterase, CER1, and serine/threonine pathways. RNA-seq analysis of leaf tissue (YL), with the exception of zinc finger, did not reveal any signal of differential gene expression.
Pathway enrichment analyses at 1 DAI and 10 DAI revealed six enriched KEGG pathways; phenylpropanoid biosynthesis (ath00940), protein processing in endoplasmic reticulum (ath04141), plant hormone signal transduction (ath04075), and plant-pathogen interaction (ath04626). Within these pathways, multiple gene groups were differentially expressed with HSP90, HSP70, HSP40, small heat shock factors (sHSFs), AUX/IAA, and cassava calmodulin-like proteins (CaM/CML) showing the most pronounced expression patterns.
Heat shock proteins HSP90, HSP70, HSP40 and sHSFs are implicated in protein processing in the endoplasmic reticulum (ER). In the resistant DSC 167, at 1 and 5 DAI, these genes were upregulated in stem and at 1 DAI in YL tissues but remained unaffected in root tissues. In contrast, HSP genes were downregulated in all tissues of DSC 260. In TMS 96/0304 all HSPs, HSP90, HSP70, HSP40, and sHSF were downregulated in the roots (Figure 6) and only HSP90 was upregulated stem tissues.
AUX/IAA proteins are part of the plant hormone signal transduction pathway. In the resistant genotypes, AUX/IAA genes were upregulated already at 1 DAI in stem tissues of DSC 167 with DSC 260 responding only at 5 DAI (Figure 6). In the susceptible TMS 96/0304, this gene group was downregulated at 10 DAI in stem and root tissues.
CaM/CML proteins (calmodulin/calmodulin-like proteins) belong to the plant-pathogen interaction pathway. CaM/CML genes were upregulated at 1 DAI in stem tissues of both resistant cassava genotypes while downregulated in roots of DSC 260. The pattern of gene regulation for each genotype, DAI and tissue type is shown in Figure 6.
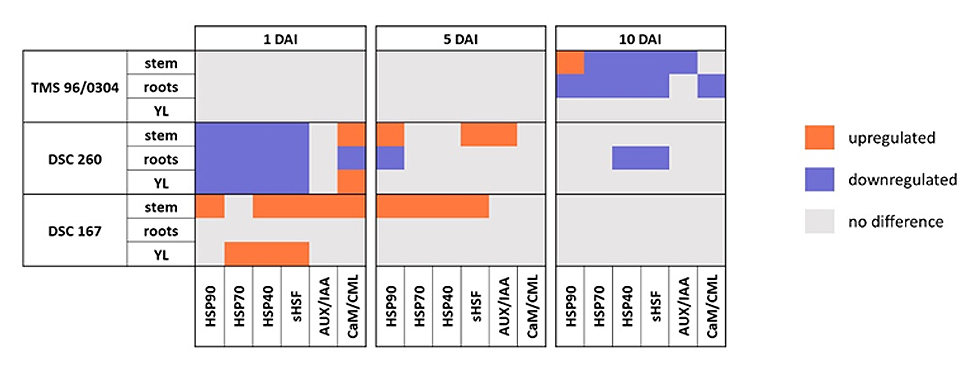 Figure 6. Heatmap of gene regulation for the three genotypes, tissue types and DAI plotted against the six gene candidates. Downregulation (blue), upregulation (orange), no difference in gene expression (grey). Significant downregulation of the HSPs in stem, roots and YL at 1 DAI in DSC 260 and upregulation of genes in tissue of the stem and young leaves of the highly resistant cassava line DSC 167. A strong differential response in the susceptible TMS 96/0304 is only visible at 10 DAI. DAI = days after inoculation; YL, young leaf; TMS 96/0304, susceptible; DSC 260, root-restricted; DSC 167, resistant.
Figure 6. Heatmap of gene regulation for the three genotypes, tissue types and DAI plotted against the six gene candidates. Downregulation (blue), upregulation (orange), no difference in gene expression (grey). Significant downregulation of the HSPs in stem, roots and YL at 1 DAI in DSC 260 and upregulation of genes in tissue of the stem and young leaves of the highly resistant cassava line DSC 167. A strong differential response in the susceptible TMS 96/0304 is only visible at 10 DAI. DAI = days after inoculation; YL, young leaf; TMS 96/0304, susceptible; DSC 260, root-restricted; DSC 167, resistant.
A temporal expression pattern plot of stem, root, and YL tissues (Figure S5) confirmed the upregulation of HSP90, HSP70, HSP40, and sHSF in stem tissue of DSC 167 at 1 and 5 DAI and in the roots at 10 DAI in contrast to the downregulation of these genes in root tissues of DSC 260 and TMS 96/0304. The gene groups were represented by multiple genes (Table 2). For HSP90, the heat shock protein 83 was found while for HSP70 the heat shock cognate 70 kDa protein 2 was identified. The DnaJ protein homolog was represented by the HSP40 in this study. Multiple sHSF-encoding genes were found in the genotypes and tissues; however, the most frequent gene was the 23.6 kDa heat shock protein. For the AUX/IAA genes three proteins of the auxin-responsive proteins were found: IAA9, IAA14, and IAA27. The CaM/CML genes were associated with multiple different calcium-binding proteins: CML23, CML41, and CML43 (Table 2). The fold changes and gene IDs are shown in Table S5.
To identify more specific transcriptional responses during early stages of CBSV-Mo83 infection, a two-factor DESeq2 analysis including genotype and time/tissue effects was performed. As already shown in Figure 4, at 1 DAI, the resistant genotypes DSC 260 and DSC 167 displayed a strong transcriptional response with most upregulated DEGs in stem and root tissues (Figure 7A). In contrast, at 10 DAI, the resistant genotypes showed a low number of mostly downregulated DEGs while at this time point a high number of DEG were found associated with the infection of TMS 96/0304 (Figure 7B). A comparison of the two resistant genotypes assigned upregulated genes to DSC 167 compared with DSC 260.
The functional annotation of DEGs from contrasting genotypes showed a high proportion of uncharacterized proteins. Several stress- and defense-related gene families were repeatedly detected across the contrasted genotypes, including WRKY transcription factors, ethylene-related genes, heat shock proteins (HSPs), auxin-related genes, cell wall–associated genes (e.g., xyloglucan-related/expansins), bHLH factors, WAT1, E3 ubiquitin ligases, serine/threonine kinases, and LRR receptor-like proteins. A 23.6 kDa small heat shock protein (mitochondrial) was upregulated in stem at 1 DAI in DSC 167 relative to DSC 260. In roots at 10 DAI, DSC 167 showed upregulation of HSP83 (HSP90), HSC70 (HSP70), and a DnaJ (HSP40) homolog compared to DSC 260. Detailed tables (protein IDs, log2FC, adjusted p-values, and annotations) are provided in Table S9, and DEG counts and direction of regulation (Table S10)
The three-factor DESeq2 likelihood ratio test (LRT) identified genotype-associated differentially expressed genes (protein IDs) in CBSV-Mo83–infected samples (Table S11). At 1 DAI, the resistant genotypes DSC 260 and DSC 167 showed more upregulated candidates in stem and roots compared with the susceptible TMS 96/0304 (Figure 8A). In DSC 260, a higher number of downregulated candidates was detected in stem and root tissues at 1 and 10 DAI, whereas DSC 167 showed more upregulated candidates in roots at 10 DAI (Figure 8B). Functional annotation indicated broadly similar protein/gene families to the standard and two-factor DESeq2 analyses, including a high proportion of uncharacterized proteins, and no clear tissue- or time-specific enrichment relative to those analyses.
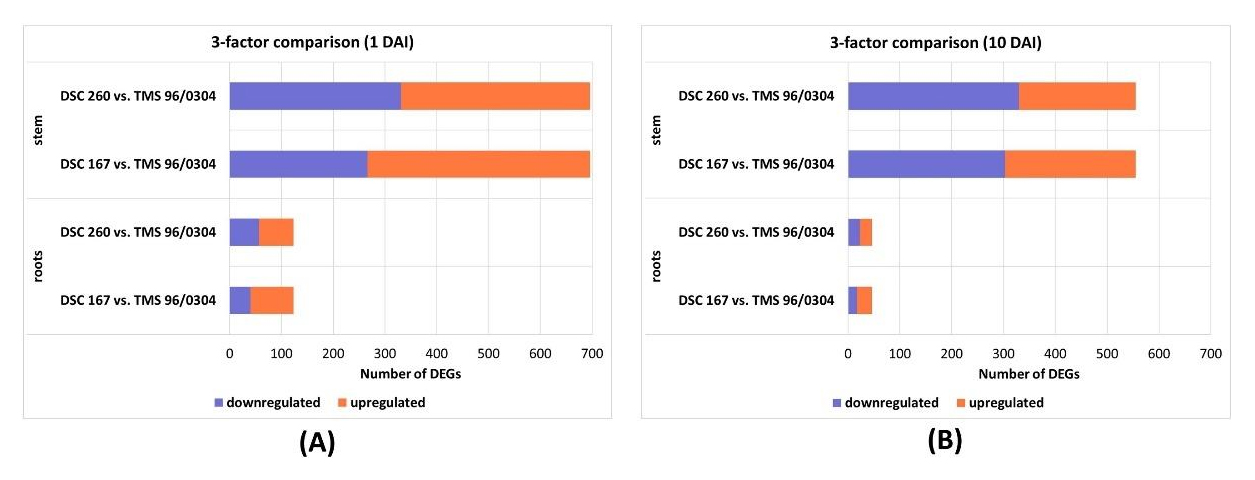 Figure 8. Bar charts of differentially expressed genes of the three-factor compariso. (A) 3 factor comparison after at 1DAI; (B) 3 factor comparison at 10 DAI; DEGs differentially expressed genes; TMS 96/0304, susceptible; DSC 260, root-restricted; DSC 167, resistant.DAI, days after inoculation; DEGs differentially expressed genes; TMS 96/0304, susceptible; DSC 260, root-restricted; DSC 167, resistant.
Figure 8. Bar charts of differentially expressed genes of the three-factor compariso. (A) 3 factor comparison after at 1DAI; (B) 3 factor comparison at 10 DAI; DEGs differentially expressed genes; TMS 96/0304, susceptible; DSC 260, root-restricted; DSC 167, resistant.DAI, days after inoculation; DEGs differentially expressed genes; TMS 96/0304, susceptible; DSC 260, root-restricted; DSC 167, resistant.
This study employed RNA-seq to investigate transcriptome responses of South American cassava genotypes with distinct resistance responses against cassava brown streak virus. By profiling gene expression at multiple time points after graft-inoculation, we aimed to uncover early molecular mechanisms that underpin resistance or susceptibility to CBSV. Our results provide a comprehensive overview of tissue specific and genotype-dependent transcriptional reprogramming, highlighting key defense-related pathways activated in resistant cultivars.
Earlier transcriptomic studies have largely focused on African cassava genotypes challenged with UCBSV and/or CBSV such as Namikonga, Kaleso, and KBH2006/18 [15–17,31]. In our study, we selected two South American cassava germplasm lines that were stringently screened for resistance against CBSV for many years and this provided the basis for our study to resolve early, tissue-specific host responses against virus infection in resistant genotypes and in the susceptible TMS 96/0304.
One of the main challenges in cassava virus research lies in the difficulties to define the onset and to achieve uniform infections. These challenges introduce variability in controlled experiments and complicate interpretation.
At this early stage, it is impossible to define the onset of CBSV infection in cassava and the invasion of tissues in individual plants of a test group. While active, CBSV replication escapes sensitive measurements in early days of infection however, as shown at 1 DAI, the response of resistant plants is highly sensitive and instant. This early time point, guided by earlier research, was chosen to capture immediate host reprogramming at or near the graft interaction zone and even before virus is detectable. The identification of viral RNA reads in the RNA-seq library at 10 DAI (Table 1) confirms viral molecules in the samples tissue but the low number of reads does not (yet) confirm a productive systemic infection even in the susceptible TMS 96/0304 but rather indicate virus invasion. Detection of CBSV reads in the resistant genotype DSC 167 at 10 DAI (Table 1 group A only) may indicate a high infection pressure from virus infected buds and virus entry followed by transport through the phloem to reach the root tissue but not progressing into a systemic infection. Indeed, detection of CBSV in DSC 167 (Table 1) can be considered virus invasion rather than proof for a uniform systemic infection.
The highly resistant genotype DSC 167 showed robust early induction of defense-related genes and the temporal structure is consistent with a rapid activation of PTI/ETI-associated modules. At 1 DAI—particularly in stem tissues, the primary site of viral entry—we observed early upregulation of receptor-like kinases/leucine-rich repeat (LRR) receptors, serine/threonine kinases, and calcium signaling components (CaM/CML), indicative of rapid pathogen perception and PTI-like signal initiation. In parallel, DSC 167 displayed strong early regulation of ER protein processing and chaperone networks, including HSP90, HSP70, HSP40/DnaJ, and small HSPs [32,33]. This is notable because these chaperone systems support immune receptor maturation and stabilization, thereby enabling effective downstream signaling. Indeed, HSP90 and HSP70 are central players in the antiviral response and together with DnaJ co-chaperones, they form complexes involved in protein folding, receptor stabilization, and immune signaling [34,35]. HSP90’s role in stabilizing NB-LRR immune receptors is well-documented in other plant-pathogen systems, including resistance against Tobacco Mosaic Virus (TMV) in Nicotiana tabacum and Arabidopsis thaliana [36–39]. In our study, strong early upregulation of HSP90 in stem tissue of DSC 167 suggests its involvement in rapid immune receptor activation and signaling and is consistent with an efficient prevention of virus infection and establishment.
Downstream of these early signaling and protein-homeostasis responses, we detected transcriptional reprogramming and defense outputs involving WRKY/MYB/NAC transcription factors, hormone-linked regulators (ethylene, jasmonate, and auxin/IAA-related genes), and structural defenses (cell wall-associated genes and phenylpropanoid biosynthesis), supporting a coordinated defense program capable of limiting virus spread. AUX/IAA and CaM/CML genes were also induced early in resistant genotypes, highlighting hormone crosstalk and calcium-mediated stress signaling as early components of defense [40,41].
In contrast to DSC 167, DSC 260 and the susceptible TMS 96/0304 exhibited showed a pronounced downregulation of immune-associated genes in root tissues at later time points, including HSPs, E3 ubiquitin ligases, aquaporins, and zinc finger transcription factors. In contrast to DSC 167, CBSV replicates in in all tissues of TMS 96/0304 and restricted to the vascular tissue, in stems and roots of DSC 260 [14] Thus, the downregulation of immune-associated genes likely facilitates viral accumulation and systemic movement. The strong response of DSC 260 at 1 DAI resembles that of TMS 96/0304 at 10 DAI (Figure 4) albeit at a lower degree of DEGs. It may indicate that immune-associated genes are downregulated to enable virus invasion in a resistant host that however restricts virus replication to the phloem companion cells of roots and stems.
The two-factor DESeq2 analysis confirmed the results obtained from the initial analysis. The high number of DEGs in stem tissue (virus entry sites) confirms early response against CBSV involving vascular tissues. The three-factor model largely recapitulated the functional trends seen in the simpler models, to highlight a large portion of uncharacterized genes implying that an increased model complexity sharpens genotype contrasts rather than revealing new tissue- or time-specific functional categories. Overall, our results support a framework in which resistance is strongly influenced by timing and localized defense activation. Rapid responses at the virus entry interface in highly resistant DSC 167 cassava versus a repression of immune response genes particularly in roots of DSC 260 (at 1 DAI) and in TMS 95/0304 (at 10 DAI) associated with virus replication. This does not explain the restriction of CBSV to the vascular tissues of stems and roots in the resistant cassava genotype DSC 260.
Our study focused on early stages of cassava infections to identify pathways and genes rather than providing functional proof. Next, candidate genes identified here will subjected to virus-induced gene silencing (VIGS) assays for functional characterization and validation. The likely best bet candidates belonging to HSP90/HSP70/HSP40 networks, calcium signaling (CaM/CML) and auxin-linked regulators (AUX/IAA). have been identified in this study.
This study identified a set of highly differentially expressed genes at early stages of CBSV infection in cassava genotypes with resistance and susceptible responses. Already at 1 DAI, the resistant lines (DSC 167 and DSC 260) responded with coordinated transcriptional changes across the phenylpropanoid pathway, ER protein processing, hormone signaling, and plant–pathogen interaction pathways, whereas the susceptible TMS 96/0304 showed a pronounced delayed response evident only at 10 DAI. Across tissues, chaperone modules (HSP90/HSP70/HSP40/DnaJ), auxin-linked regulators (AUX/IAA; IAA9/14/27), and calcium signaling components (CaM/CML; CML23/41/43) consistently associate with resistant phenotypes. Taken together, our results support the notion that rapid activation of immune-response genes close to viral entry sites prevent virus replication in DSC 167. While virus replication takes place, the early activation of genes in DSC 260 prevents plant invasion and restricts the virus to the phloem companion cells of the roots.
The following supplementary materials are available online, Figure S1–S4: PCA plots of different cassava tissues E, Figure S5: Temporal expression pattern plots of protein candidates based on the log2FC at 1, 5, and 10 days after inoculation; Table S1: Sampling strategy; Table S2: qRT-PCR results for replicates sampled; Table S3: Raw reads, summary and mapping of stem samples; Table S4: Raw reads, summary and mapping of root samples; Table S5: Raw reads, summary and mapping of YL samples; Table S6: Raw reads, summary and mapping of OL samples; Table S7: Numbers of DEGs; Table S8: List of gene names and functions.; Table S9: Protein ID, log2 fold change, adjusted p-value, and protein function of DEGs after two-factor DESeq2 analyses; Table S10: Number DEGs_DESeq2_2factor_LRT_3factor; Table S11: Protein ID, log2 fold change of DSC 260, log2 fold change of DSC 167, adjusted p-value, and protein function of DEGs after three-factor DESeq2-LRT analyses.
The RNA-seq data that support the findings of this study have been deposited into European Nucleotide Archive (ENA): PRJEB88994.
Conceptualization, SS and SW; methodology, SS, SW, JL and BP; validation, SS; formal analysis, JL, SS; investigation, JL, SS; writing—review and editing, JL, SW, BP and SS; supervision, SS and BP; project administration, SS and SW; funding acquisition, SW. All authors have read and agreed to the published version of the manuscript.
The authors declare no conflicts of interest.
This research project was funded by the One CGIAR Initiative Research Program on Roots, Tubers, and Ba-nanas (CRP-RTB) through the International Institute of Tropical Agriculture, grant number ID INV-041105 (BMGF-CIP).
We are grateful for the assistance during the experiments provided by the team of the Plant Virus Department of the DSMZ. This work was supported by the de.NBI Cloud within the German Network for Bioinformatics Infrastructure (de.NBI) and ELIXIR-DE (Forschungszentrum Jülich and W-de.NBI-001, W-de.NBI-004, W-de.NBI-008, W-de.NBI-010, W-de.NBI-013, W-de.NBI-014, W-de.NBI-016, W-de.NBI-022). We also want to thank Google Colab for help to create the heatmap and the bar chart of the rRNA check.
1.
2.
3.
4.
5.
6.
7.
8.
9.
10.
11.
12.
13.
14.
15.
16.
17.
18.
19.
20.

Copyright © Hapres Co., Ltd. Privacy Policy | Terms and Conditions